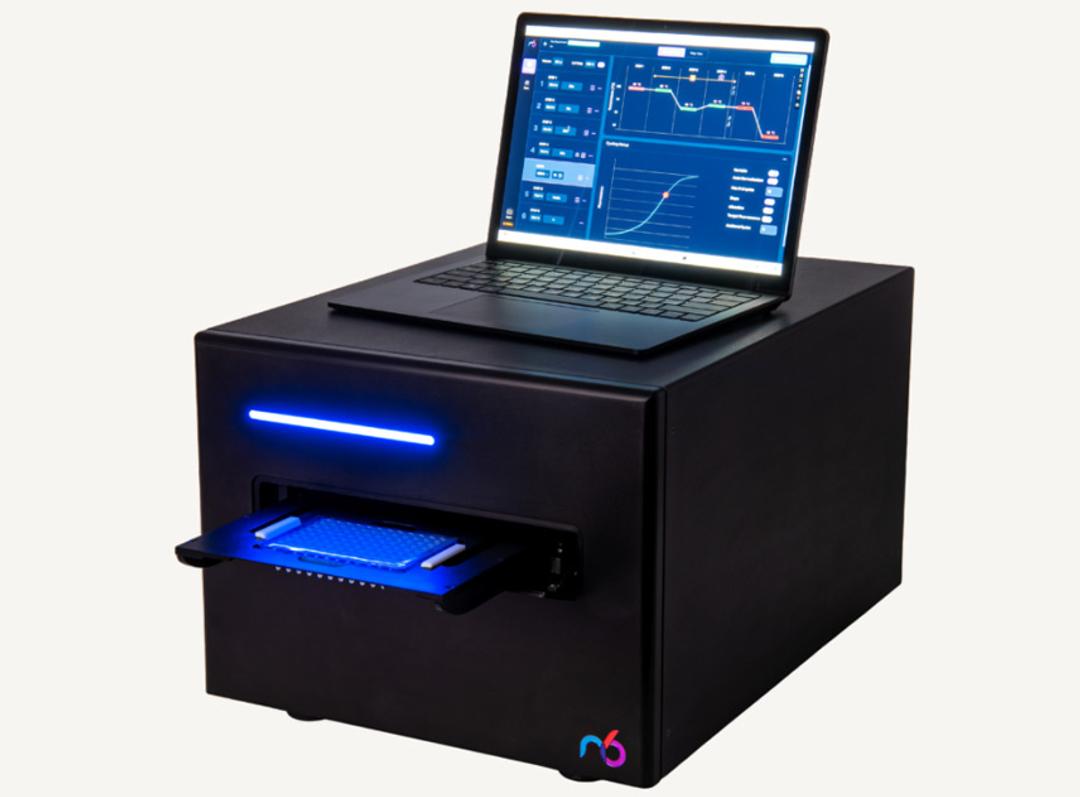
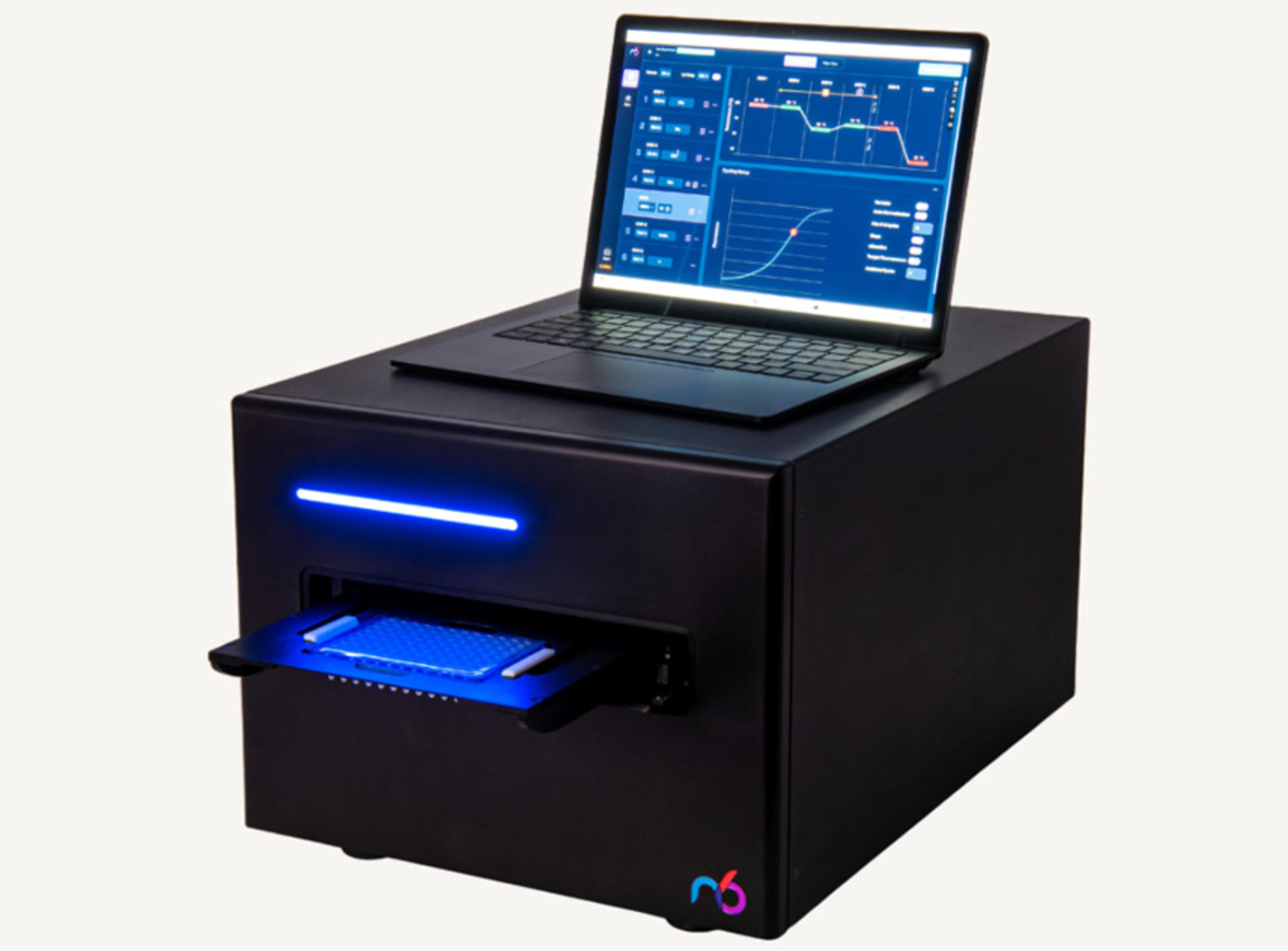
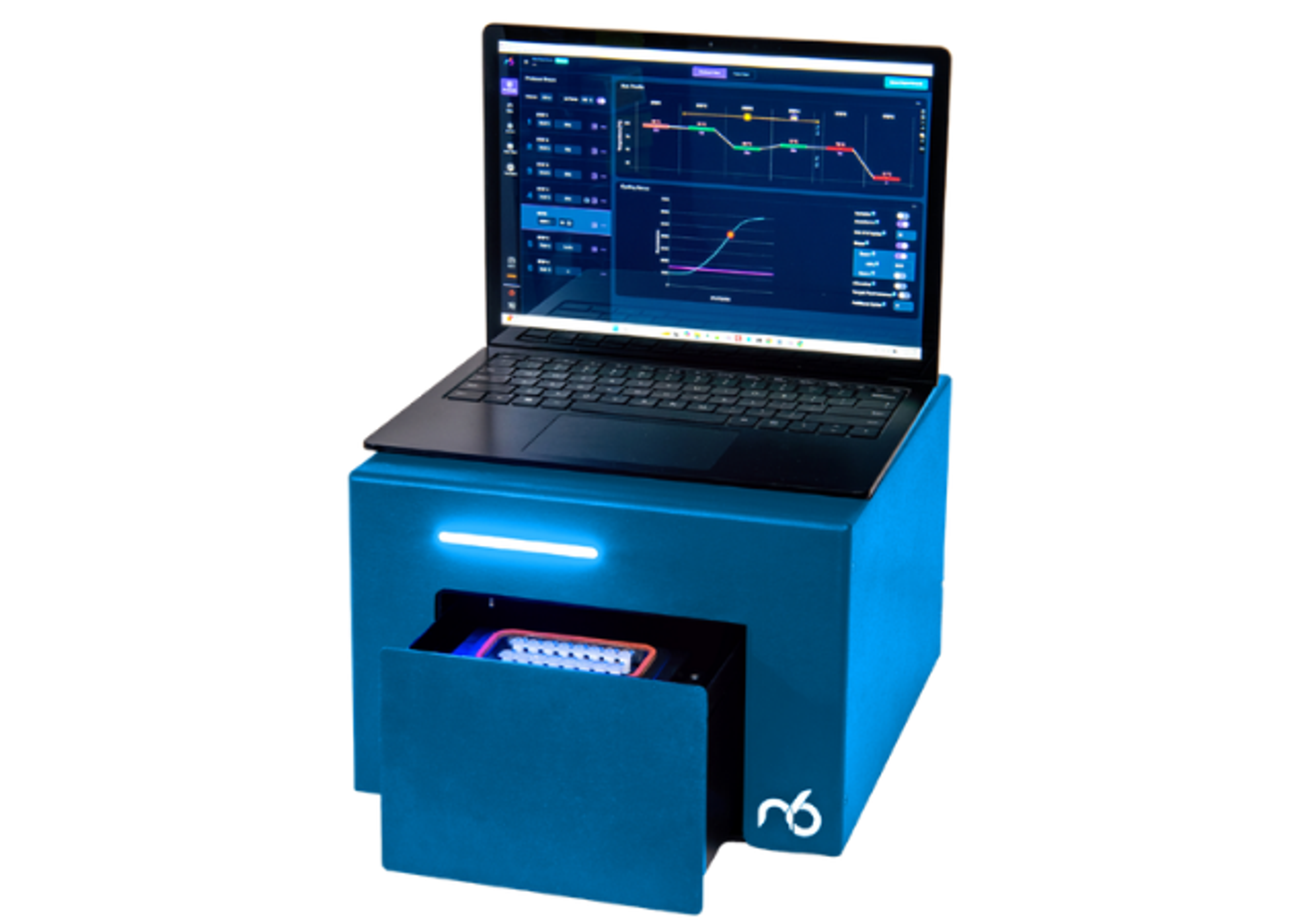
icon96™
Stop gambling on cycle numbers. The icon96 uses iconPCR™ technology with AutoNorm™ software to provide real-time fluorescence feedback in each of 96 independent wells, preventing under-amplification dropouts and over-amplification artifacts—delivering balanced, sequencing-ready libraries without manual quantification or normalization.
Every NGS library prep risks under-amplifying precious samples into dropouts or over-amplifying them into chimera-filled garbage data—and you won't know until it's too late. The icon96 solves this with iconPCR™ technology: 96 individually controlled thermal zones with real-time fluorescence detection. AutoNorm™ software monitors every well and stops each reaction at its optimal cycle, ensuring data quality and workflow efficiency in one automated step.
Improve Sequencing Data Quality:
- Eliminate dropouts: Real-time monitoring prevents under-amplification failures—critical for precious clinical samples like FFPE, cfDNA, or biopsies you can't re-collect
- Reduce artifacts: AutoNorm stops reactions before over-cycling creates PCR duplicates, chimeras, or adapter dimers that compromise sequencing data quality
- Rescue variable-quality samples: Automatically adjusts cycles for degraded or low-input samples (DIN 1.3–8), avoiding both failed libraries and over-amplification distortions
Relieve Library Prep Workflow Bottlenecks:
- 96 → 1 cleanup: Pool libraries by volume directly after PCR; perform one SPRI cleanup instead of 96 (saves ~$225/plate)
- No manual normalization: AutoNorm delivers every sample at the same target fluorescence—sequence-ready without Qubit quantification, TapeStation analysis, or dilution steps
- Flexible throughput: Run RNA-seq, 16S metagenomics, single-cell, WGS, or FFPE workflows (and more coming) on the same instrument with any existing kit—no proprietary consumables required
- Time savings: Complete library prep to sequencing pool in under 3 hours versus 8+ hours with traditional workflows
How It Works:
The icon96 features 96 miniaturized thermocyclers with precision temperature control and real-time fluorescence detection in every well (iconPCR technology). AutoNorm monitors library amplification every cycle and automatically stops individual wells when they reach optimal yield—preventing both under- and over-amplification across variable inputs (0.6 ng to 1000 ng) without prior sample quantification.
The Bottom Line - Why it's iconic:
- 96 wells, each with independent control
- AutoNorm: No more manual quant or normalization
- Every sample gets its perfect cycling—no batching
- Real-time feedback, automatic cycle adjustment
- Save low-input and tough samples with fewer failures
- One simple cleanup—ditch individual SPRIs
- Works with your kits and workflows—no “locked-in” extras
- Faster prep, lower costs, better data quality
Brochures
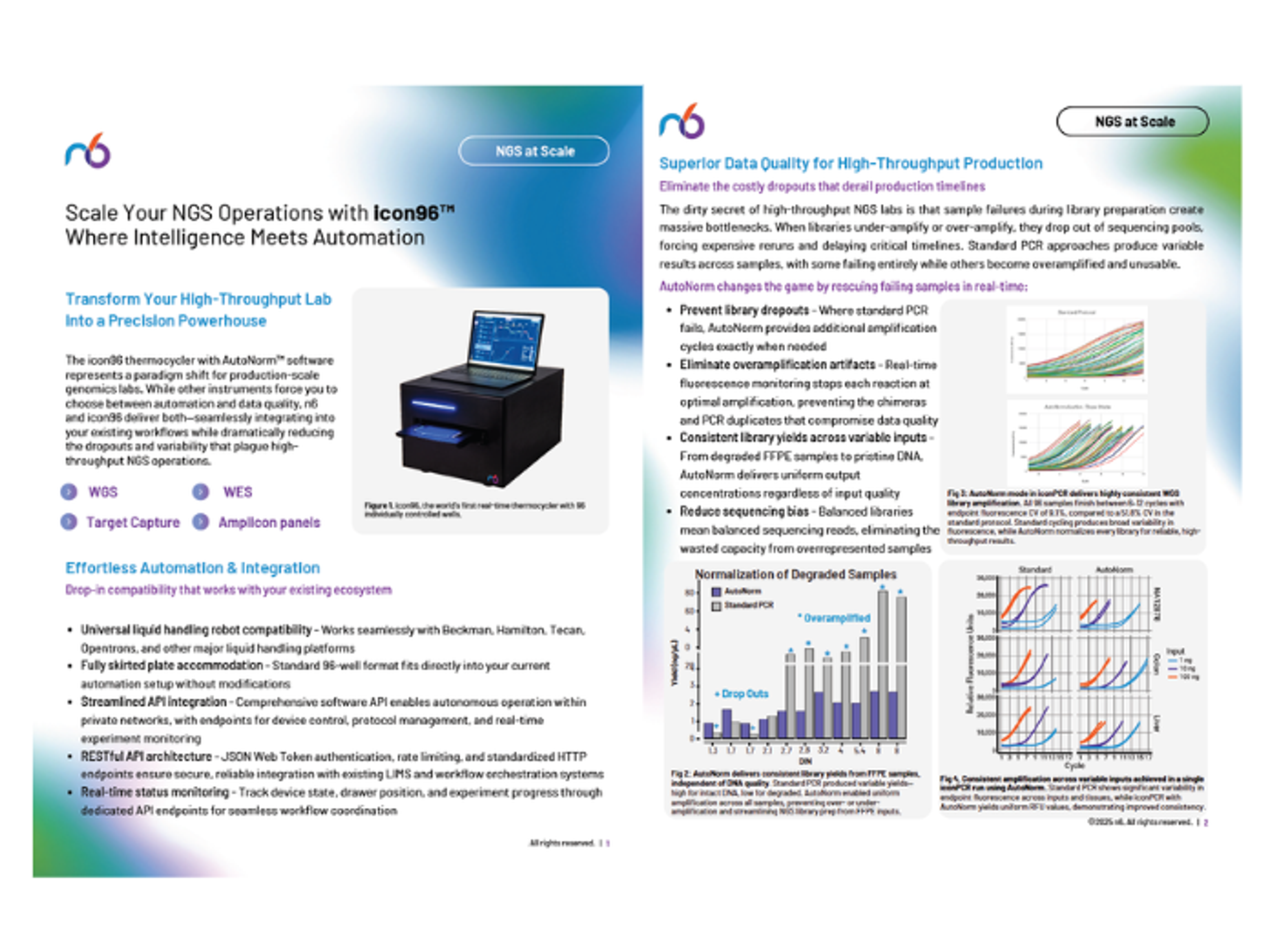
Scale your NGS operations with icon96
Find out how to transform your high-throughput lab into a precision powerhouse with the icon96 thermocycler with AutoNorm™ software from n6. This solution promises both automation and data quality, seamlessly integrating into your existing workflows while reducing the dropouts and variability that plague high-throughput NGS operations.
icon96 - Amplify, quantify, and normalize all at once
Learn more about iconPCR™ from n6, the world’s first thermocycler with 96 individually controlled wells and built-in AutoNorm™ software, designed for normalized, sequencing-ready libraries in one step.
Sensitive cfDNA sequencing using adaptive PCR control with icon96
This application note describes how adaptive, real‑time PCR amplification using the icon96™ thermocycler with AutoNorm™ technology enables highly sensitive sequencing of cell‑free DNA (cfDNA) from ultra‑low input samples. Developed in collaboration with the Dana‑Farber Cancer Institute, the workflow demonstrates uniform library yields across variable inputs, improved on‑target capture efficiency, and reliable detection of variants down to 0.5% allele frequency, even from just 1 ng of cfDNA.
Balanced sequencing pools through controlled amplification using AutoNorm™
This application note demonstrates how iconPCR™ technology with AutoNorm™ on the icon96 real-time thermocycler enables balanced sequencing pools by dynamically controlling PCR amplification in each well. Unlike conventional fixed-cycle PCR workflows that require post-PCR quantification and normalization, AutoNorm monitors fluorescence in real time and stops amplification once a target threshold is reached. This automated normalization produces consistent library yields across variable inputs, simplifies NGS library preparation, reduces hands-on time by up to 60%, minimizes failed libraries, and maintains balanced read distribution for whole genome sequencing.
Power up your assay development: DOE made easy on icon96™
Design of Experiments (DOE) can dramatically improve PCR assay development, but traditional thermocyclers limit flexibility and efficiency. The icon96™ platform overcomes these limitations with 96 independently controlled wells, enabling multiple thermocycling conditions to be tested in a single plate. This technical note demonstrates how icon96 simplifies DOE workflows, reduces experimental runs, and accelerates assay optimization while minimizing time and consumable usage.
AutoNorm™ of NEXTFLEX small RNA libraries using the iconPCR™ system
Explore how Revvity’s NEXTFLEX Small RNA-Seq Kit v4, combined with n6’s iconPCR system and AutoNorm technology, enables adaptive, real-time normalization of small RNA libraries from diverse human tissues, reducing PCR cycles and variability in library yield without altering RNA composition profiles.
Unlocking superior microbial profiling with iconPCR
16S rRNA gene sequencing is essential for exploring microbial diversity in various ecosystems, including soil, water, and human associated microbiomes. However, conventional PCR methods used in 16S library preparation often introduce significant biases due to overamplification, under-amplification, and the generation of chimeric sequences.
Discover how the icon96™, developed by n6, represents a transformative shift in PCR technology. Summarized in this comparative study, learn how using icon96 with iconPCR and conventional PCR workflows can produce high data quality, taxonomic resolution, and workflow consistency, especially when paired with long-read platforms like PacBio for full-length amplicon sequencing.
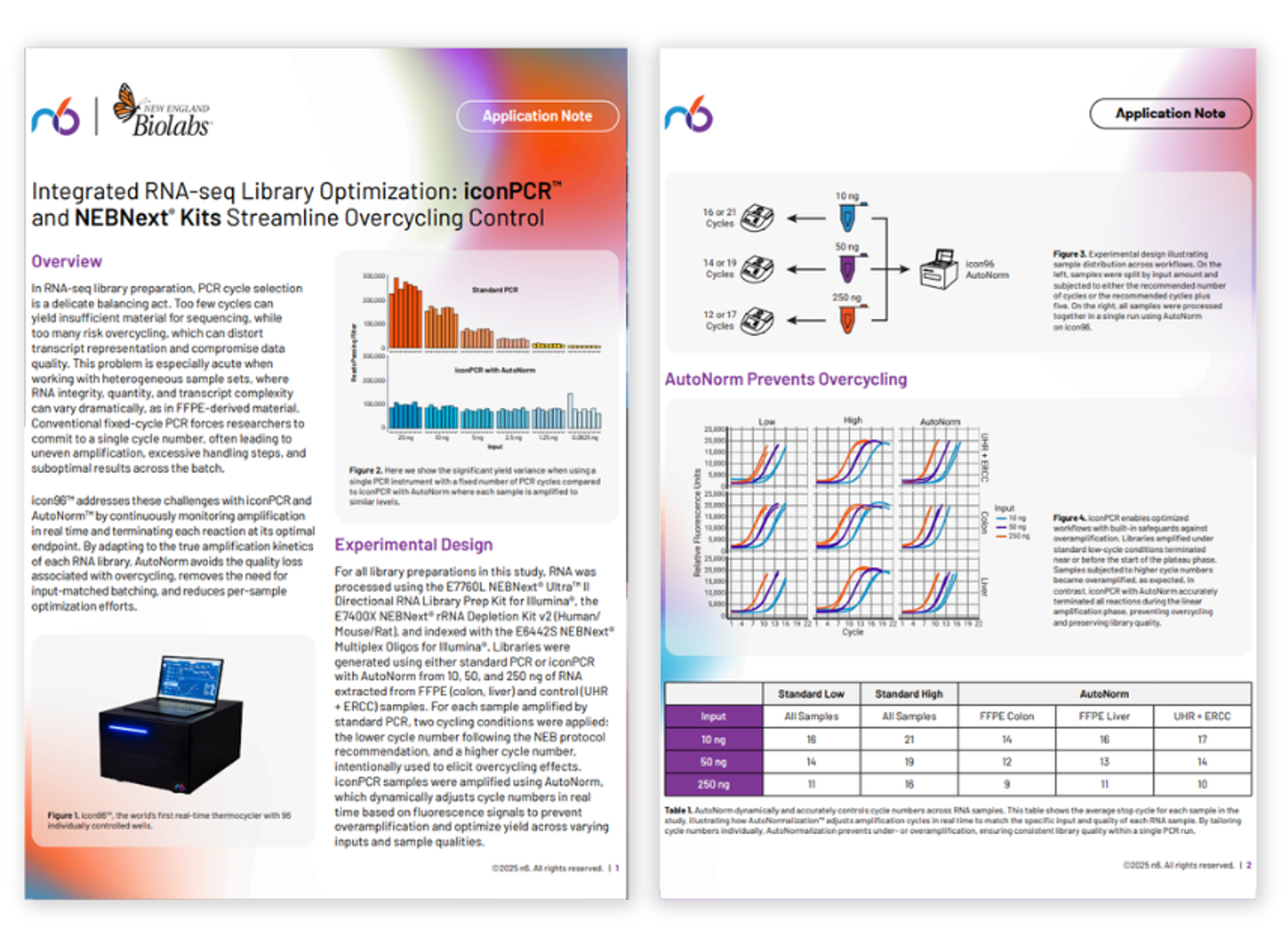
Integrated RNA-seq library optimization with iconPCR and NEBNext kits
Single‑cell workflows are highly sensitive to sample quality, handling steps, and instrument settings, making it challenging to generate reliable, biologically meaningful data. Variability introduced during dissociation, staining, or capture, can reduce cell viability, skew population representation, and compromise downstream analysis.
This resource explores practical strategies to optimize each stage of single‑cell preparation, helping researchers improve sample integrity, reduce technical noise, and increase data quality. Learn how to gain clear guidance to streamline workflows, strengthen reproducibility, and achieve more confident single‑cell insights.
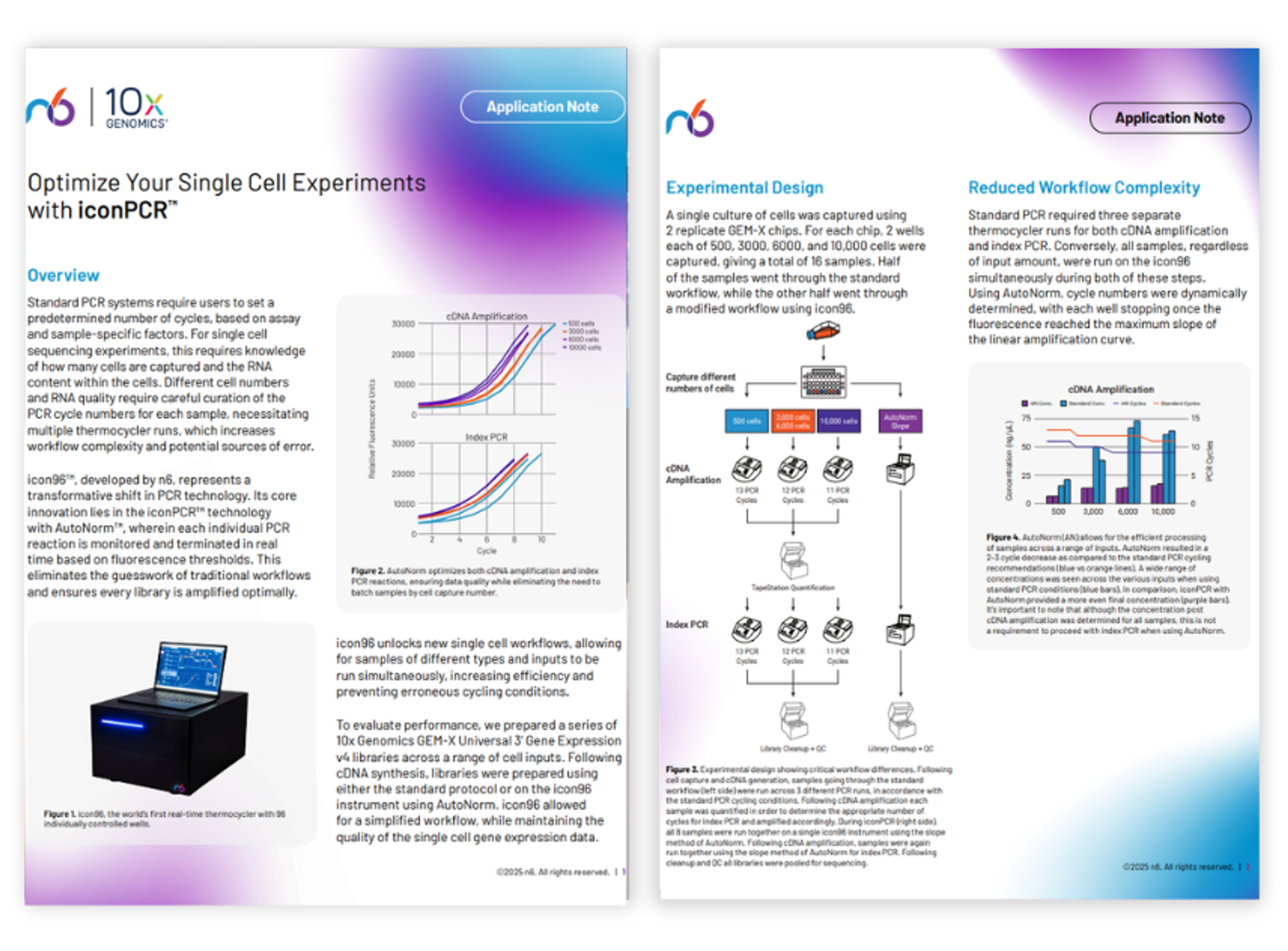
Optimize your single cell experiments with iconPCR
Standard PCR systems require users to set a predetermined number of cycles, based on assay and sample-specific factors. For single cell sequencing experiments, this requires knowledge of how many cells are captured and the RNA content within the cells. Different cell numbers and RNA quality require careful curation of the PCR cycle numbers for each sample, necessitating multiple thermocycler runs, which increases workflow complexity and potential sources of error.
Discover how the icon96™, when used together with AutoNorm™ can monitor each individual PCR reaction and terminate in real time based on fluorescence thresholds. This eliminates the guesswork of traditional workflows and ensures every library is amplified optimally.
Streamlined FFPE DNA library prep using NEBNext UltraShear and Multiplex Oligos kits with iconPCR
n6 Tec and NEB offer a streamlined FFPE DNA library preparation workflow that enhances amplification consistency, reduces hands-on time by up to 60%, and maintains high data quality across variable inputs.
This application note highlights an optimized workflow using NEBNext UltraShear and NEBNext Multiplex Oligos with iconPCR AutoNorm, which dynamically tunes PCR cycling, minimizes batching and QC burden, and delivers uniform, high-quality NGS libraries from challenging FFPE samples.
How to optimize low-input cfDNA library prep for targeted sequencing sensitivity
Wednesday, March 25 at 15:00 GMT | 16:00 CET | 11:00 EDT | 8:00 PDT
What if your PCR step could adapt to every sample, instead of forcing every sample to adapt to your PCR?
In this now-on-demand webinar, Zach Herbert from Dana-Farber Cancer Institute's Molecular Biology Core Facilities shares how his team developed a targeted cfDNA sequencing workflow capable of detecting variants at <0.5% allele frequency from as little as 1 ng input — equivalent to just ~300 genome equivalents.
Watch this SelectScience® webinar to see real data from a genomics core lab:
- Library yield CV dropped from 54% to 8% (10 ng input) — 6x more uniform than fixed-cycle PCR
- On-target rates jumped from ~70% to >90% after implementing sample-specific amplification, across 1 ng and 10 ng inputs
- ~100 patient-derived cfDNA samples (2–25 ng range) processed in a single run, delivering balanced read counts (50–100M reads/pool) for hybrid capture
- Mean target coverage >200x at 1 ng input, with reliable variant detection down to 0.5% VAF using duplex sequencing
The challenge solved: cfDNA workflows are molecule-limited. Fixed-cycle PCR either over-amplifies (artifacts, distorted duplex families) or under-amplifies (insufficient yield for capture). The result? Imbalanced pools, wasted sequencing budget, and compromised sensitivity for rare variants.
The solution: Real-time, per-well PCR control that terminates amplification at each sample's optimal endpoint, preserving molecules, ensuring sufficient yield, and enabling mixed-input runs (1 ng + 10 ng together).
Why this matters for your lab:
"icon96 is really a game changer — shining a light into that black box [of PCR]. We can now see exactly the amplification path of every sample."
— Zach Herbert, Dana-Farber Cancer Institute
90%+ on-target isn't a benchmark. It's what Dana-Farber achieved, and what your cfDNA workflow can achieve.
Certificate of attendance
If you attend the live webinar, you will automatically receive a certificate of attendance, including a learning outcomes summary, for continuing education purposes.
If you view the on-demand webinar, you can request a certificate of attendance by emailing editor@selectscience.net.
Webinar details
- Cost: Free to attend
- Location: Online
- Duration: 60 minutes
Registration is required to secure your place. If you register but can’t attend live, you will receive a link to the on‑demand recording once it becomes available.
Scale genomics without compromising data integrity on icon96
One of the biggest challenges in scaling genomics is maintaining data quality as throughput increases. As applications like single-cell analysis, spatial biology, MRD, cancer diagnostics, and microbiome research become more complex, traditional PCR and normalization workflows are struggling to keep up. In this SelectScience interview, Pranav Patel, Founder and CEO of n6 Tec, explains how the team is tackling this challenge with icon96™, the company's flagship amplification technology. By intelligently adjusting amplification based on sample quality and quantity, icon96 preserves data integrity, improves reproducibility, and eliminates the need for brute-force normalization.
This video was filmed at SLAS 2026.