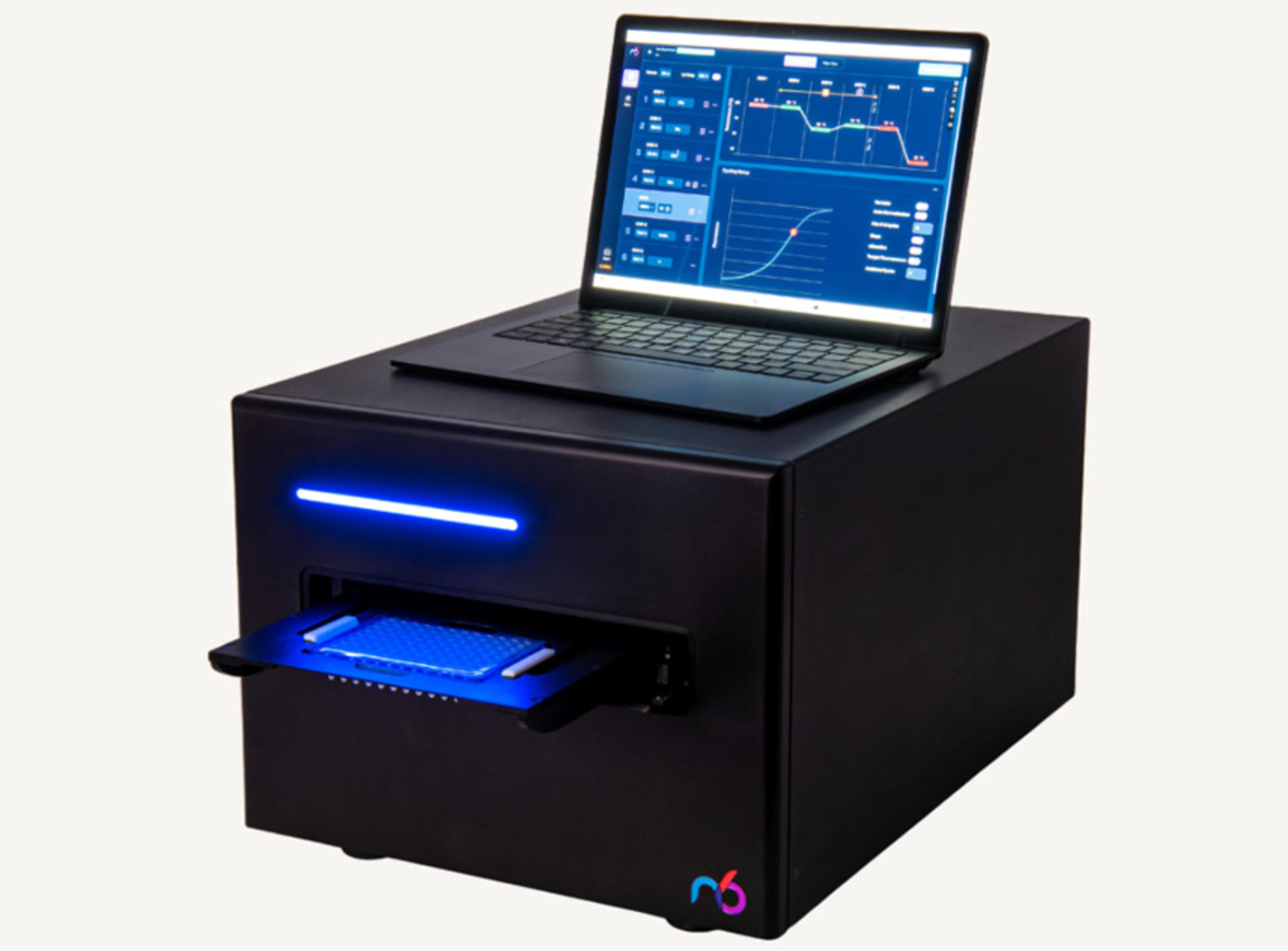
How to optimize low-input cfDNA library prep for targeted sequencing sensitivity
Wednesday, March 25 at 15:00 GMT | 16:00 CET | 11:00 EDT | 8:00 PDT
What if your PCR step could adapt to every sample, instead of forcing every sample to adapt to your PCR?
In this now-on-demand webinar, Zach Herbert from Dana-Farber Cancer Institute's Molecular Biology Core Facilities shares how his team developed a targeted cfDNA sequencing workflow capable of detecting variants at <0.5% allele frequency from as little as 1 ng input — equivalent to just ~300 genome equivalents.
Watch this SelectScience® webinar to see real data from a genomics core lab:
- Library yield CV dropped from 54% to 8% (10 ng input) — 6x more uniform than fixed-cycle PCR
- On-target rates jumped from ~70% to >90% after implementing sample-specific amplification, across 1 ng and 10 ng inputs
- ~100 patient-derived cfDNA samples (2–25 ng range) processed in a single run, delivering balanced read counts (50–100M reads/pool) for hybrid capture
- Mean target coverage >200x at 1 ng input, with reliable variant detection down to 0.5% VAF using duplex sequencing
The challenge solved: cfDNA workflows are molecule-limited. Fixed-cycle PCR either over-amplifies (artifacts, distorted duplex families) or under-amplifies (insufficient yield for capture). The result? Imbalanced pools, wasted sequencing budget, and compromised sensitivity for rare variants.
The solution: Real-time, per-well PCR control that terminates amplification at each sample's optimal endpoint, preserving molecules, ensuring sufficient yield, and enabling mixed-input runs (1 ng + 10 ng together).
Why this matters for your lab:
"icon96 is really a game changer — shining a light into that black box [of PCR]. We can now see exactly the amplification path of every sample."
— Zach Herbert, Dana-Farber Cancer Institute
90%+ on-target isn't a benchmark. It's what Dana-Farber achieved, and what your cfDNA workflow can achieve.
Certificate of attendance
If you attend the live webinar, you will automatically receive a certificate of attendance, including a learning outcomes summary, for continuing education purposes.
If you view the on-demand webinar, you can request a certificate of attendance by emailing editor@selectscience.net.
Webinar details
- Cost: Free to attend
- Location: Online
- Duration: 60 minutes
Registration is required to secure your place. If you register but can’t attend live, you will receive a link to the on‑demand recording once it becomes available.
Speakers


Moderator

Who should attend?
- Liquid biopsy / cfDNA / ctDNA workflows
- Minimal residual disease (MRD) monitoring
- Low-input targeted panels (<10 ng)
- Duplex sequencing / hybrid capture
- Precious samples (plasma, CSF, FFPE)
What will this webinar cover?
- How to achieve uniform library yields across 12x input range (2–25 ng) in a single run
- The step-by-step optimizations that drove >90% on-target rates
- Real sequencing metrics from ~100 clinical cfDNA samples
- Live Q&A insights from Dana-Farber's core facility