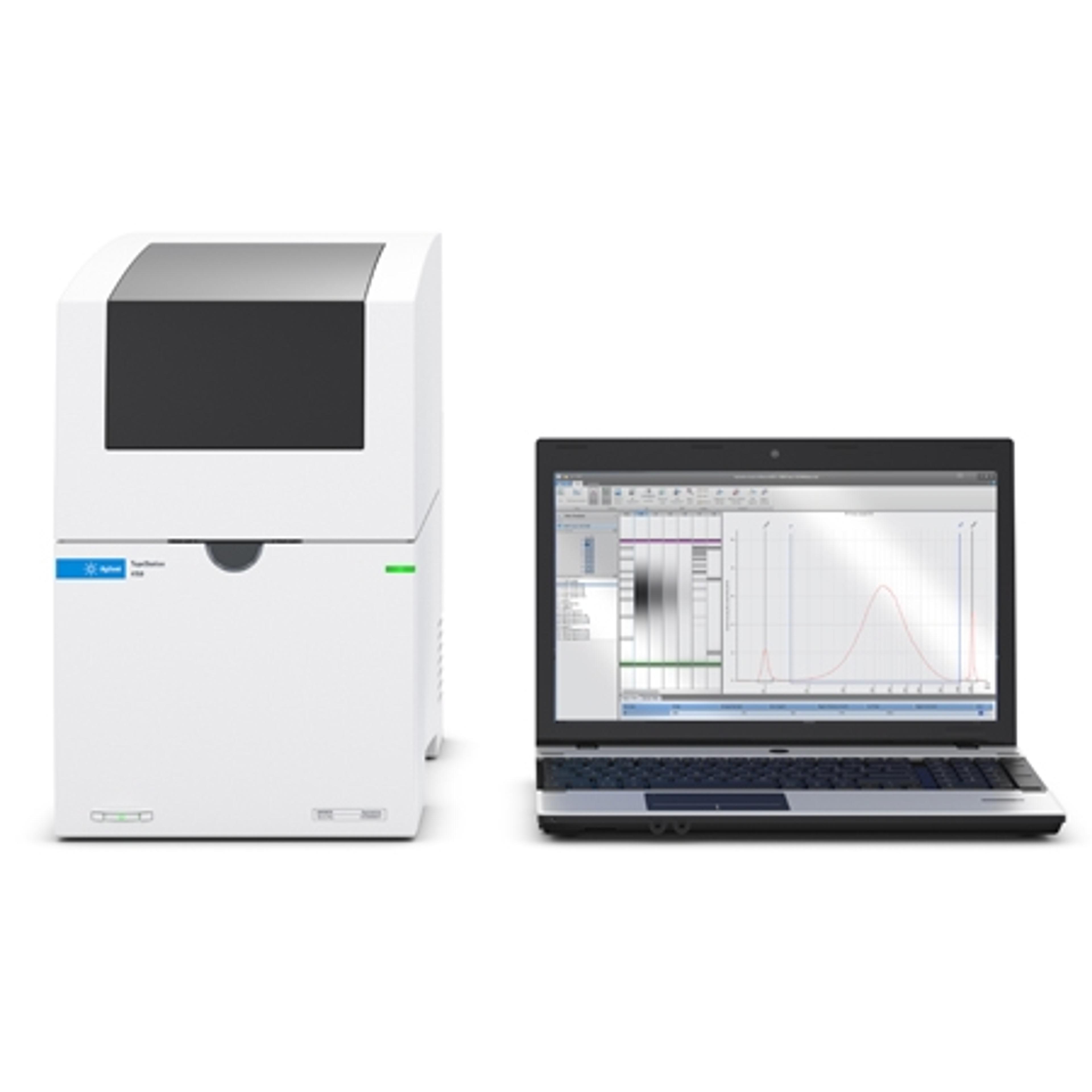
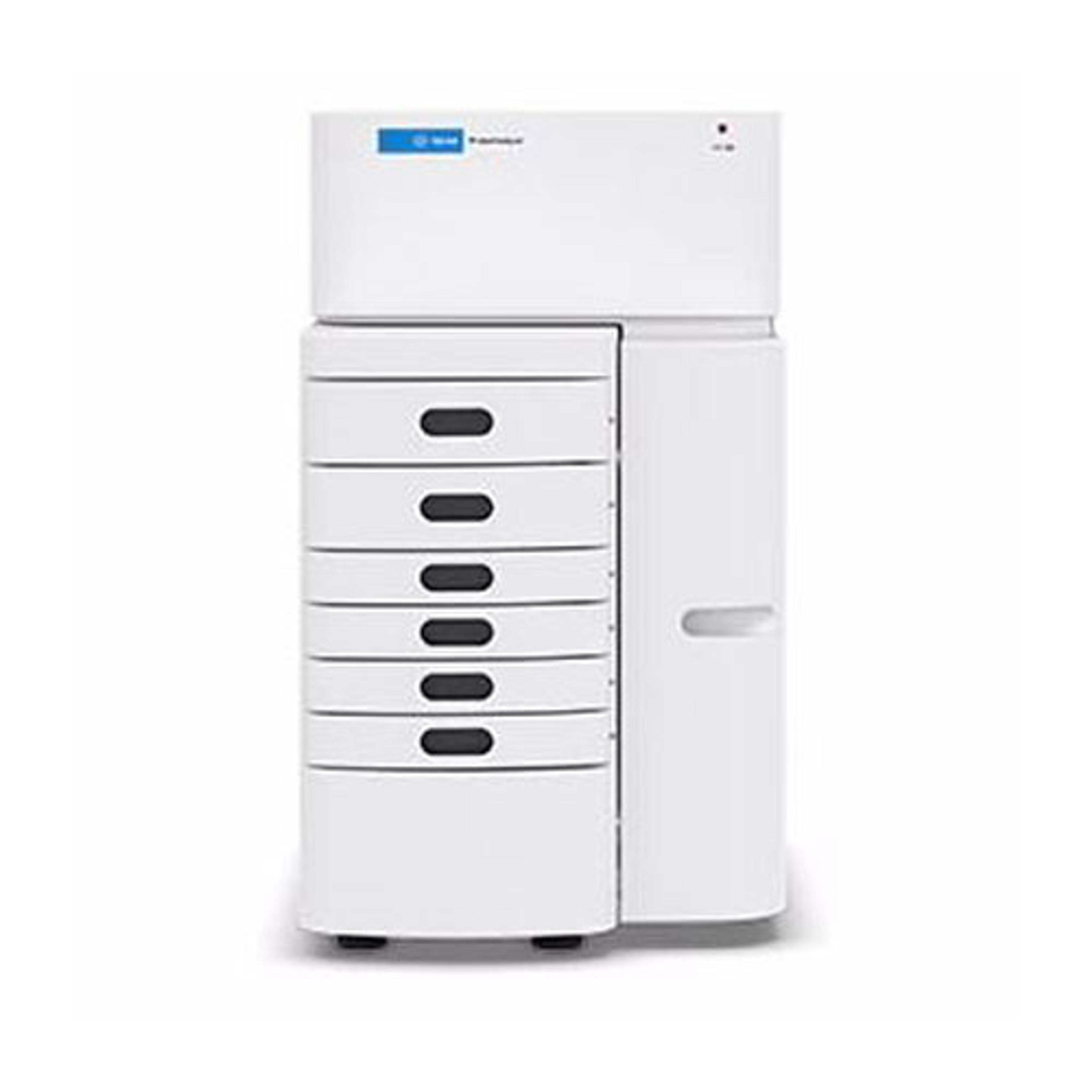
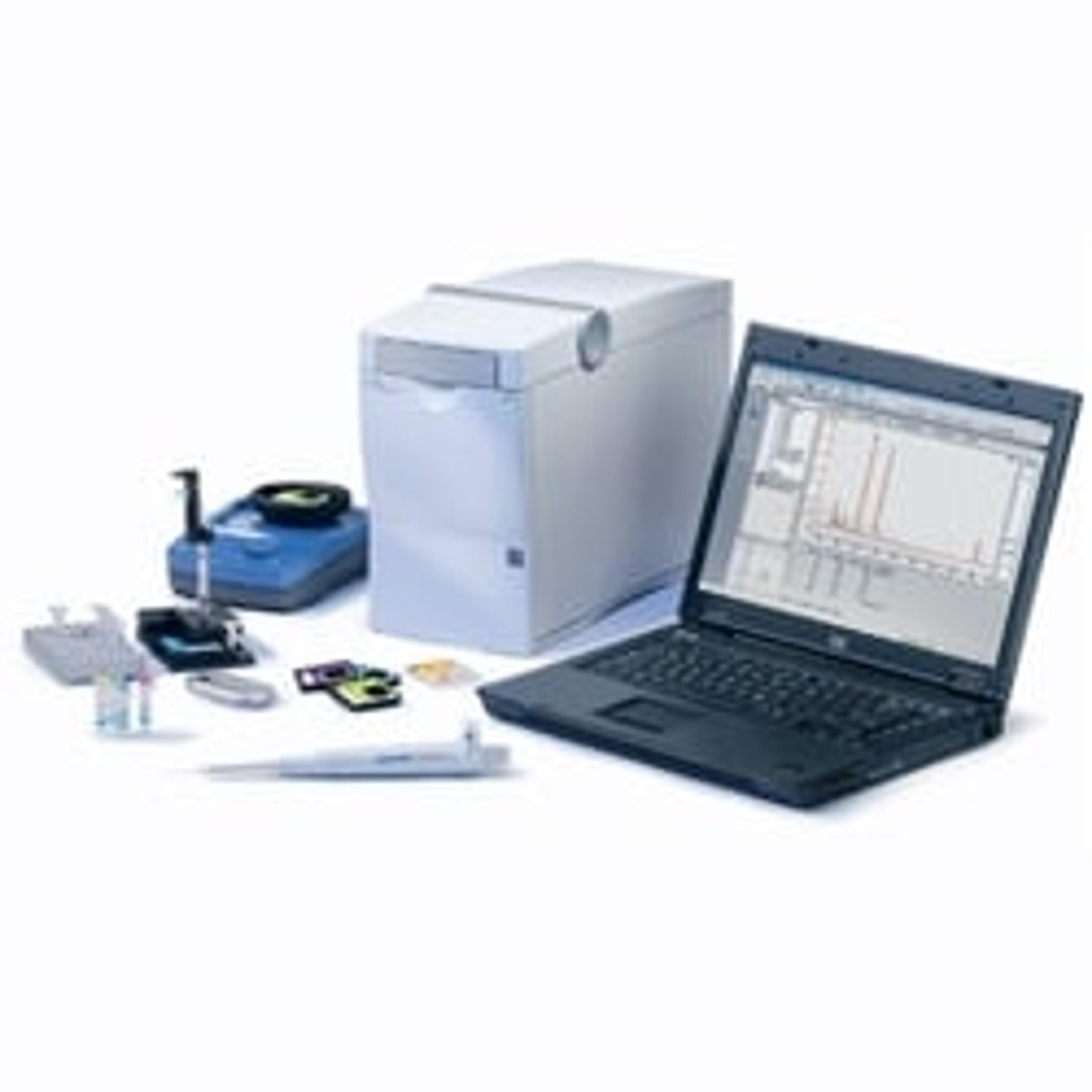
4150 TapeStation system
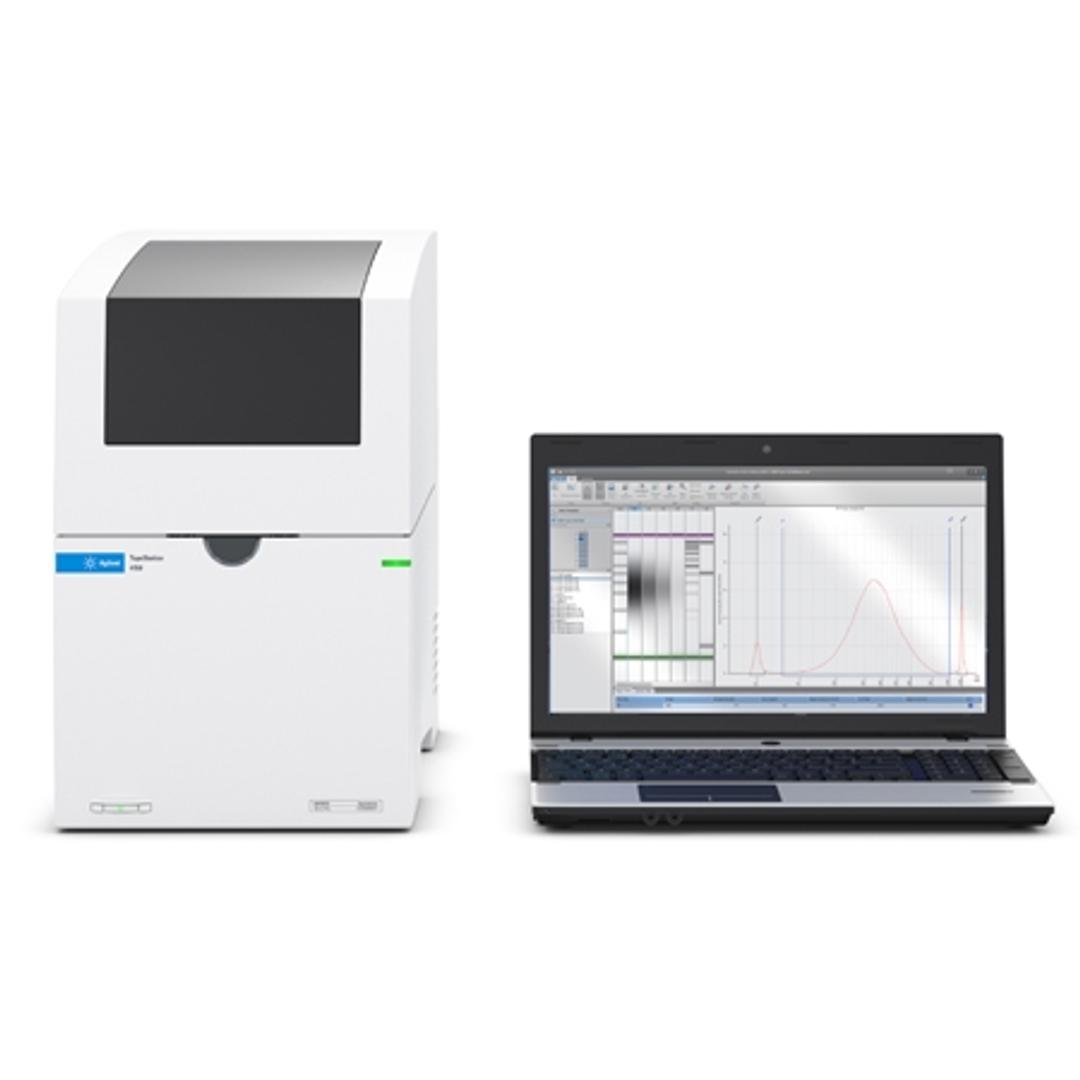
The 4150 TapeStation system is an economic, entry-level system which comes with data processing software, reagents, and ScreenTape devices for analysis of sample size, quantity, and integrity of 1 to 16 samples. Delivering highly accurate and precise analytical evaluation, the systems fit perfectly into next-generation sequencing (NGS) or biobank workflows for handling smaller numbers of samples.
The 4150 TapeStation system is an economic, entry-level system which comes with data processing software, reagents, and ScreenTape devices for analysis of sample size, quantity, and integrity of 1 to 16 samples. Delivering highly accurate and precise analytical evaluation, the systems fit perfectly into next-generation sequencing (NGS) or biobank workflows for handling smaller numbers of samples.
Brochures
Agilent automated electrophoresis portfolio brochure
Whether performing low- or ultra-high-throughput testing, Agilent Technologies' automated electrophoresis instrument portfolio delivers reliable, objective assessments of sample integrity, concentration, and fragment size. Explore the complete portfolio of automated electrophoresis instruments, application-specific assays, intuitive software, and expert technical services that provide flexible solutions for robust and objective sample quality assessment.
Quality control in the PacBio HiFi whole genome sequencing library preparation workflow
Accurate and reliable next-generation sequencing (NGS) results rely on starting with high-quality samples. In long-read sequencing workflows, such as the PacBio HiFi workflow, essential quality control (QC) steps include assessing input gDNA, intermediate steps throughout library preparation, and the final library. Electrophoretic analysis enables evaluation of sample size, purity, and integrity at each of these steps. Agilent offers several automated electrophoresis systems, including the Agilent Femto Pulse, Fragment Analyzer, and TapeStation systems, each with unique benefits for DNA QC.
This application note demonstrates the capabilities and limitations of each of the automated electrophoresis systems within the PacBio HiFi whole genome sequencing (WGS) library preparation workflow, aiding in the selection of the appropriate QC system.
NGS analysis of hereditary cancer predispositions at the Laboratory of Molecular Genetics, Jihlava Hospital
Explore how the Laboratory of Molecular Genetics at Jihlava Hospital, Czech Republic is utilizing the Agilent 4150 TapeStation system to streamline its NGS workflow, ensuring that only high-quality samples are sequenced.
The team is working to validate an NGS method designed to analyze hereditary predispositions to various cancers, including breast and colon. Their workflow involves isolating DNA from blood and buccal swab samples, with DNA concentration measured post-isolation with a fluorometer. Discover how the lab uses the TapeStation system at two stages, during library prep and before sequencing to assess the quality and quantity of their libraries prior to pooling.
Performance characteristics of the RNA and the high sensitivity RNA ScreenTape assays for the 4150 TapeStation System
The Agilent 4150 TapeStation system is an automated electrophoresis solution for fast and reliable nucleic acid sample quality control. The entire Agilent RNA and DNA ScreenTape portfolio is applicable on the 4150 TapeStation system. The Agilent RNA ScreenTape assays enable separation, integrity, and quantity analysis of total RNA samples from both eukaryotic and prokaryotic origin. The RNA integrity number equivalent (RINe) provides objective assessment of RNA degradation. The 4150 TapeStation system together with the RNA ScreenTape assays is perfectly suited for RNA quality control at low throughput need. In this technical overview, Agilent Technologies focuses on the performance characteristics of the RNA ScreenTape assay and the Agilent High Sensitivity RNA ScreenTape assay (HS RNA ScreenTape assay) on the 4150 TapeStation system regarding RNA integrity analysis, sensitivity, and quantification. The performance of the 4150 TapeStation system was compared to the 4200 TapeStation system using the same assays to demonstrate full compatibility. Furthermore, the Agilent 2100 Bioanalyzer system, together with the corresponding RNA assays as widely accepted standards for RNA quality control, were applied for a benchmarking of all three systems.
The European Reference Genome Atlas initiative: A collaborative effort to advance our understanding of European biodiversity
Reference genomes are a powerful method to gain complete insight into the biology of unique species within our communities. They are critical for understanding the function of biodiversity in protecting threatened species, a concern of scientists worldwide. Considering this, researchers from across Europe decided to collaborate and form the European Reference Genome Atlas (ERGA) initiative. ERGA is a pan-European consortium of multidisciplinary scientists dedicated to generating a high-quality, complete genome for all European species, with 20 percent of those species being at risk of extinction. Here, Agilent Technologies shows how the 4150 TapeStation System is supporting this research.
Quality control in Illumina sequencing workflows using the TapeStation system
In this application note, Agilent Technologies describes quality control (QC) steps in various Illumina next-generation sequencing (NGS) workflows provided as a service by the German Cancer Research Center (DKFZ) Genomics and Proteomics Core Facility. The QC steps can be performed using the Agilent 4200 TapeStation system with the ScreenTape portfolio. Appropriate reference samples are included as positive controls to monitor the individual steps during library preparation. Representative QC data of these reference samples serve as positive examples for successful completion of critical steps of the most frequently used NGS library protocols.
Building with quality: Incorporating NGS approaches to infectious disease, pathology, and oncology research in Latvia
In this case study, Agilent Technologies shows how a laboratory team in Latvia were able to successfully manage the transition from Sanger sequencing-based approaches to next-generation sequencing (NGS). Here, they share the insights they gained throughout the successful adoption of the new workflow.
Automated nucleic acid sample QC in NGS workflows
High-quality DNA and RNA samples support strong next-generation sequencing (NGS) results. To help prevent suboptimal sequencing data and avoid wasting time and resources, quality control (QC) steps are recommended at various points in the NGS workflow, from initial sample assessment to final library qualification for NGS library preparation protocols. Here, Agilent Technologies explains how the benefits of the Agilent TapeStation systems and ScreenTape technology offer ideal and easy-to-use sample QC solutions for NGS workflows.
FFPE sample quality control for the MGISEQ-2000 sequencing platform with the Agilent TapeStation system
Formalin-fixed paraffin embedding (FFPE) is one of the more common ways to preserve clinical samples, and FFPE tumor samples serve as valuable study materials for clinical and translational medicine research. In this application note, learn more about the Agilent 4150 TapeStation system. This new technology was used to conduct sample QC of an entire library construction process for five FFPE DNA samples used on the MGI tech MGISEQ-2000RS sequencing platform. Explore how high-quality sequencing results were achieved, and more.
Quality matters: Tackling high molecular weight DNA contamination in cell-free DNA for optimal liquid biopsy NGS
Tuesday, June 24, 2025 at 16:00 BST / 17:00 CEST / 11:00 EDT / 08:00 PDT
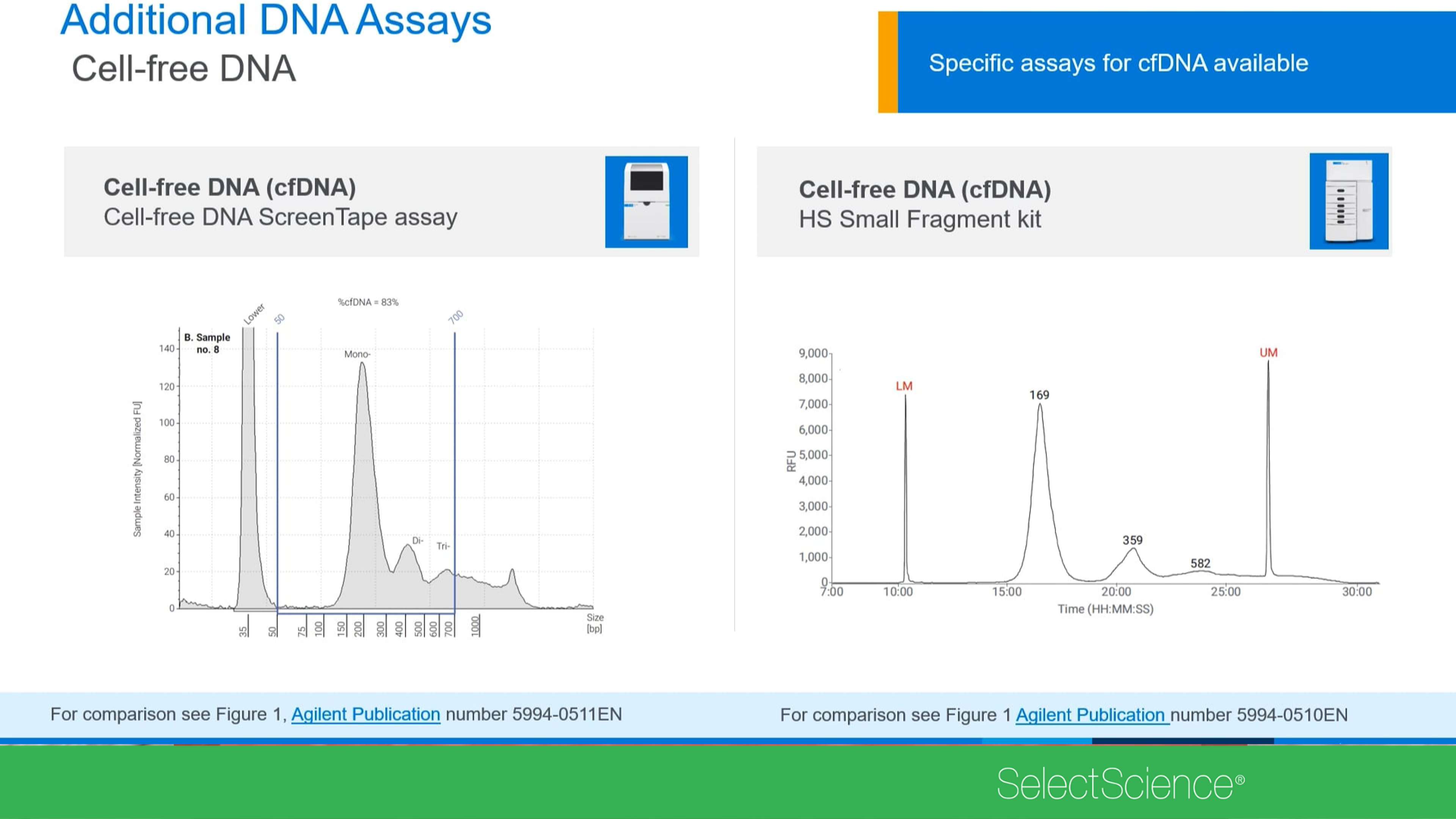
Cell-free DNA (cfDNA) has become an essential tool in clinical research and serves as valuable biomarker in liquid biopsy applications. However, cfDNA presents a challenge for Next-Generation Sequencing (NGS) workflows due to its low yield, complex fragmentation pattern and potential presence of contaminating high molecular weight (HMW) DNA.
In this webinar, our expert speaker will discuss reliable pre-analytical quality control using the Agilent Cell-free DNA ScreenTape assay and the TapeStation systems, emphasizing their relevance to NGS-based liquid biopsy applications.
Key learning objectives
- Understand the challenges associated with cfDNA extraction and HMW DNA contamination.
- Learn about the impact of HMW DNA on cfDNA quality and NGS results.
- Explore how to ensure pre-analytical quality control of cfDNA samples.
- Discover the unique benefits of the Agilent Cell-free DNA ScreenTape assay for obtaining the full picture of your cfDNA quality.
Who should attend?
NGS and genomics researchers, molecular biologists, researchers in cancer diagnostics and liquid biopsy, and sequencing lab managers and technicians
Certificate of attendance
If you attend the live webinar, you will automatically receive a certificate of attendance, including a learning outcomes summary, for continuing education purposes.
If you view the on-demand webinar, you can request a certificate of attendance by emailing editor@selectscience.net.
For Research Use Only. Not for use in diagnostic procedures. PR7001-4363
How automation and integrated sample QC enhance a targeted long-read-sequencing workflow
Tuesday, November 4, at 16:00 GMT | 17:00 CET | 11:00 EST | 8:00 PST
As the demand for greater accuracy and efficiency in genetic analysis grows, new approaches are needed to overcome the blind spots of conventional sequencing methods. This webinar will showcase a fully automatable workflow that integrates Agilent SureSelect hybrid capture, Agilent automated electrophoresis systems for sample quality control with PacBio HiFi long-read sequencing.
In addition to discussing workflow, we’ll share case studies where long reads play a fundamental role in identifying genetic variants that may be missed by conventional sequencing in research studies.
Join us to learn how targeted long-read sequencing:
- Focuses on genes and regions of interest in genetic research, while reducing costs and boosting sequencing depth at key loci.
- Overcomes the limitations of short-read technologies, including challenges with pseudogenes, homologous loci, structural variants, complex transcript isoforms, splicing events, and difficult regions such as repetitive or GC-rich sequences.
- Enhances both analytical performance and laboratory efficiency through automation.
Key learning objectives:
- Gain practical insights into implementing an automated targeted long-read sequencing workflow in the laboratory.
- Identify the genetic mutation types best suited for targeted long-read sequencing approaches.
- Understand the analytical advantages of long-read sequencing compared with conventional short-read methods.
Who should attend?
Lab directors, molecular pathologists, research scientists, clinical lab technicians, and anyone running or interested in running NGS workflows.
Certificate of attendance
If you attend the live webinar, you will automatically receive a certificate of attendance, including a learning outcomes summary, for continuing education purposes.
If you view the on-demand webinar, you can request a certificate of attendance by emailing editor@selectscience.net.
For Research Use Only. Not for use in diagnostic procedures. PR7003-294
Achieve successful sequencing data based on input sample and library quality control
Next-generation sequencing (NGS) has transformed the way genomes are analyzed. As technologies trend towards higher throughput, lower cost, and improved data quality, standardization and quality control remain key topics in the NGS workflow.
In this SelectScience NGS webinar series, produced in partnership with Agilent Technologies, we explore the importance of input sample quality control (QC). The webinar demonstrates how sample QC relates to the quality of downstream sequencing results and examines key QC aspects to consider when scaling NGS throughput and automation. This series features presentations from expert speakers with a variety of scientific backgrounds and consists of two webinars, starting on Friday, 6 May 2022.
Achieve successful sequencing data based on input sample and library quality control
This webinar will demonstrate how rigorous quality control (QC) of samples and libraries is crucial to producing high-quality data in next-generation sequencing (NGS). Sample QC metrics like concentration, DNA and RNA integrity numbers, 28S/18S ratio, DV200, fragment length, and amount of adapter dimers will be covered. How these metrics provide reasonable estimates for downstream NGS sequencing parameters such as on-target rate, duplication rate, and sequencing coverage will be discussed.
Carsten Maus, Ph.D., is a bioinformatician at the NGS Core Facility of the German Cancer Research Center (DKFZ) in Heidelberg. Dr. Maus shares his experience on the dependencies between the quality of samples, libraries, and data within a high-throughput sequencing unit.
Rainer Nitsche, Ph.D., Product Manager, Agilent Technologies, will give a brief overview of the Agilent TapeStation systems and how they can be used to collect QC metrics from RNA and DNA samples.
Key learning objectives
- The connection between key NGS QC metrics and downstream sequencing results
- Methods to secure optimal NGS results by applying sample QC
- Advantages of the TapeStation automated electrophoresis systems
Who should attend?
- Genomics researchers using NGS
- Molecular biologists
- Cancer researchers
- Lab managers and technicians
Certificate of attendance
All webinar participants can request a certificate of attendance, including a learning outcomes summary, for continuing education purposes.
Optimization and Quality Control of Library Preparation in Bulk and Single-Cell ATAC Sequencing
Advancements in sequencing techniques have allowed for detailed exploration of chromatin accessibility, but the efficiency and quality of library preparation remain crucial to achieving reliable outcomes. This webinar delves into the optimization and quality control methodologies for the preparation of sequencing libraries in both bulk-ATAC seq and single-cell ATAC seq using the Agilent TapeStation system. Our expert speakers will discuss findings from their study and how it offers insights into improving the efficiency and reliability of ATAC seq library preparations across different scales. Find out how their team investigated and optimized various conditions, including the Tn5 enzyme treatment duration and concentration for bulk-ATAC seq, and further explore the determination of ideal lysis time for efficient nuclear isolation in the context of single-cell ATAC seq.
Key learning objectives
- Understand how integral quality control is in the ATAC-Seq workflow
- Learn how to improve the efficiency and reliability of ATAC-Seq library preparations across different scales
- Discover the importance of lysis and fragmentation conditions during ATAC-Seq library preparation
Certificate of attendance
All webinar participants can request a certificate of attendance, including a learning outcomes summary, for continuing education purposes.
Pre-Analytical Considerations for Minimal Residual Disease Detection by Next-Generation Sequencing
For liquid biopsy applications, high recovery of total cfDNA is essential for the detection of ctDNA. Due to requisite low limits of detection, accurate tumor-informed minimum residual disease (MRD) detection by next-generation sequencing (NGS) testing is highly dependent upon maximum recovery of cfDNA from plasma. Conventional cfDNA extraction methods, often subsequently adapted to liquid biopsy applications, fail to deliver sufficient yield or the analytical sensitivity required for early detection of liquid biopsy markers, resulting in assay failure (e.g. ‘QNS errors’).
In this webinar, experts from Strata Oncology will highlight the importance of cfDNA yield and purity using the Revolution Sample Prep System from nRichDx for cfDNA extraction as well as reliable pre-analytical quality control with the Agilent Cell-free DNA ScreenTape assay and the TapeStation systems, and their relation to NGS-based MRD testing.
Key learning objectives
- Learn how to gain high recovery of total cfDNA in liquid biopsy applications such as accurate tumor-informed minimum residual disease by NGS testing
- Discover the importance of high cfDNA yield and purity during cfDNA extraction
- Explore how to ensure pre-analytical quality control of cfDNA samples
Certificate of attendance
All webinar participants can request a certificate of attendance, including a learning outcomes summary, for continuing education purposes.
Ensuring quality control in next-generation sequencing
Explore the various systems from Agilent Technologies including the Bioanalyzer, TapeStation, and Fragment Analyzer, that can address challenges in next-generation sequencing (NGS) quality control. Gain top tips on how you can improve your workflow and transform your NGS applications with reliable quality metrics.
Exploring cell-free DNA and small RNA analysis techniques
Discover how you can sensitively determine sample quality using the Bioanalyzer, TapeStation and Fragment Analyzer from Agilent Technologies. These automated electrophoresis systems can be used for cell-free DNA (cfDNA) and small RNA quality control and analysis. Delivering highly accurate and precise analytical evaluation, learn how these systems fit perfectly into next-generation sequencing (NGS), biobank or vaccine development workflows for low to high sample throughput.
Sample QC control in multiple workflows
Discover Agilent’s automated electrophoresis solutions designed to help ensure experimental success through accurate sample quality assessment. With a focus on NGS and protein QC, these solutions offer flexible throughput and high-resolution performance to support and streamline your workflow.