Scorpions Primers
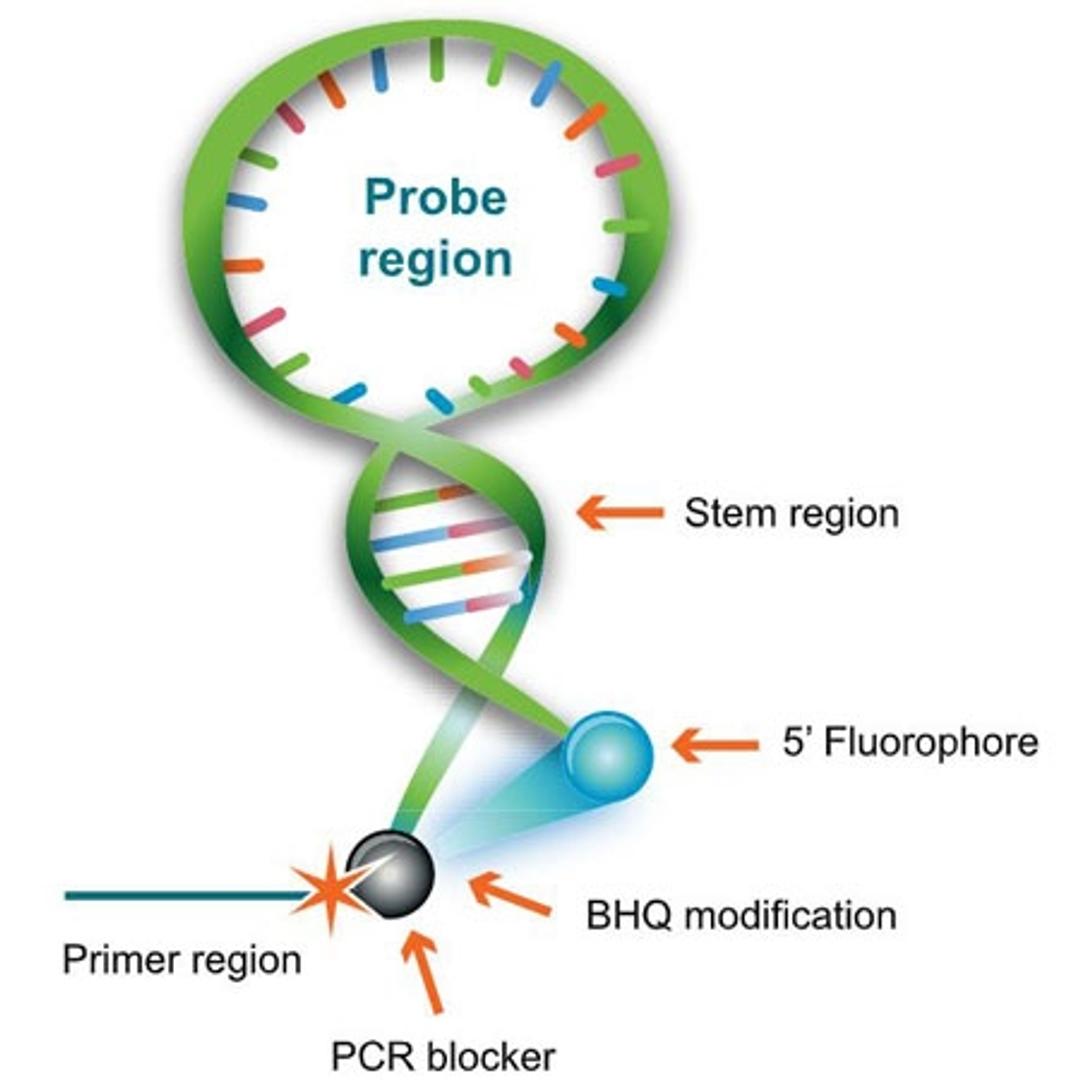
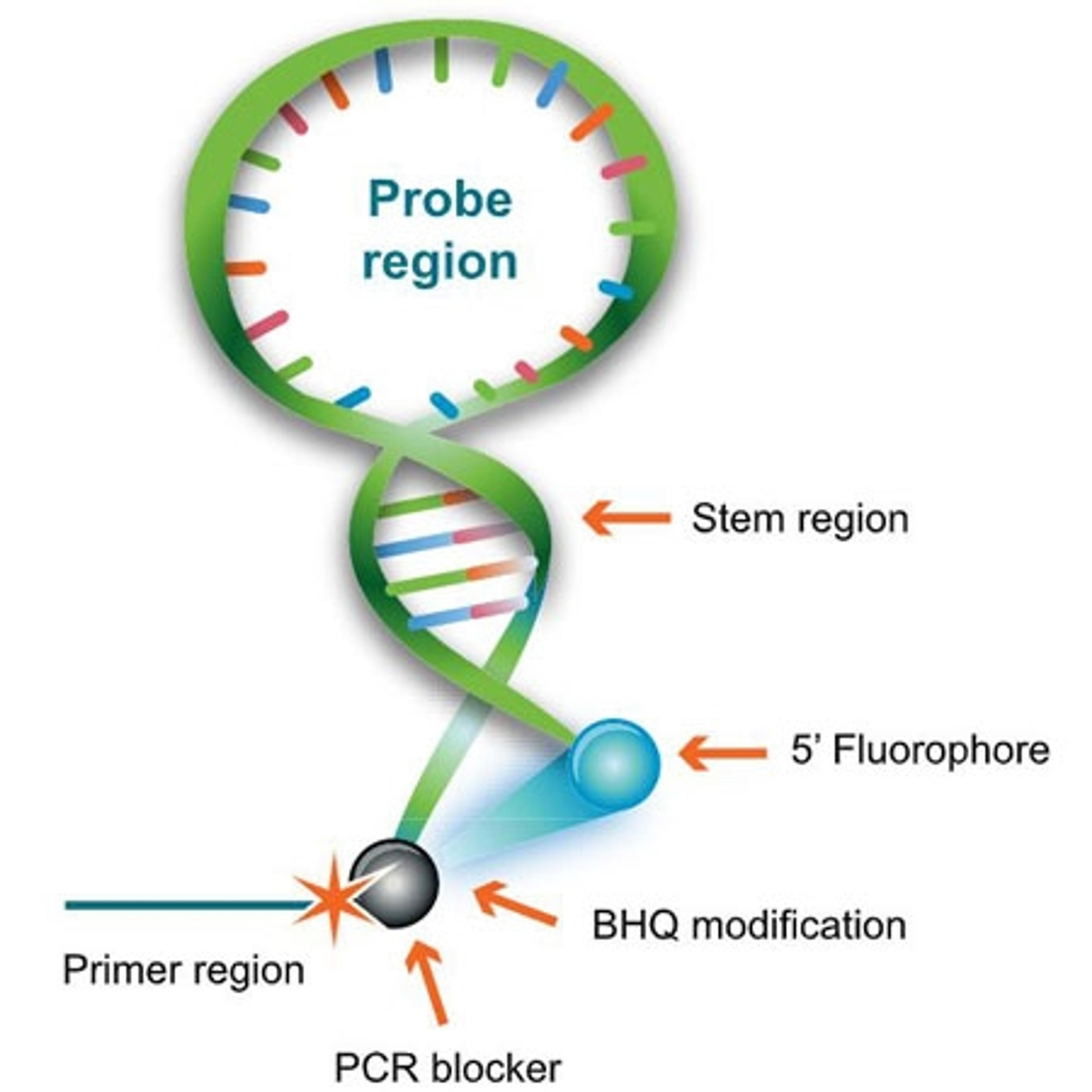
A dual-labelled probe that combines a hairpin structure and a PCR primer element in a single oligo, allowing for target detection through a unimolecular mechanism. Mismatched hybrids are less stable than the reformed stem region. Therefore, Scorpions Primers only produce signal when the probe region hybridizes to a perfect-match target sequence within the primer extension product.
BHQ™ (Black Hole Quencher™)-labeled Scorpions primers for PCR analysis are unlike dual-labeled probes and Molecular Beacons because they combine primer and probe in one molecule, with the primer at the 3' end and the probe contained within a hairpin-loop structure at the 5' end.
Scorpions primers utilize a uni-molecular mechanism that acts faster in solution for instantaneous fluorescence in real-time PCR. Similar to Molecular Beacons, these primers do not require enzymatic cleavage of the probe during PCR cycling. These qualities make Scorpions Primers valuable tools for rapid, real-time PCR, endpoint PCR, SNP detection, and gene quantification.
Scorpions primers provide a number of important advantages over other PCR Probe formats:
- Exceptional specificity
- Excellent S:N
- Easily multiplexed
Scorpions architecture
Scorpions Primers are probe/primer hybrids unlike other FRET probes, whose design is such that they emit light only when incorporated into the PCR product during amplification. These primers incorporate two distinct structures: 1) a target-specific DNA probing sequence and 2) a target-specific PCR Primer.
DNA probing sequence
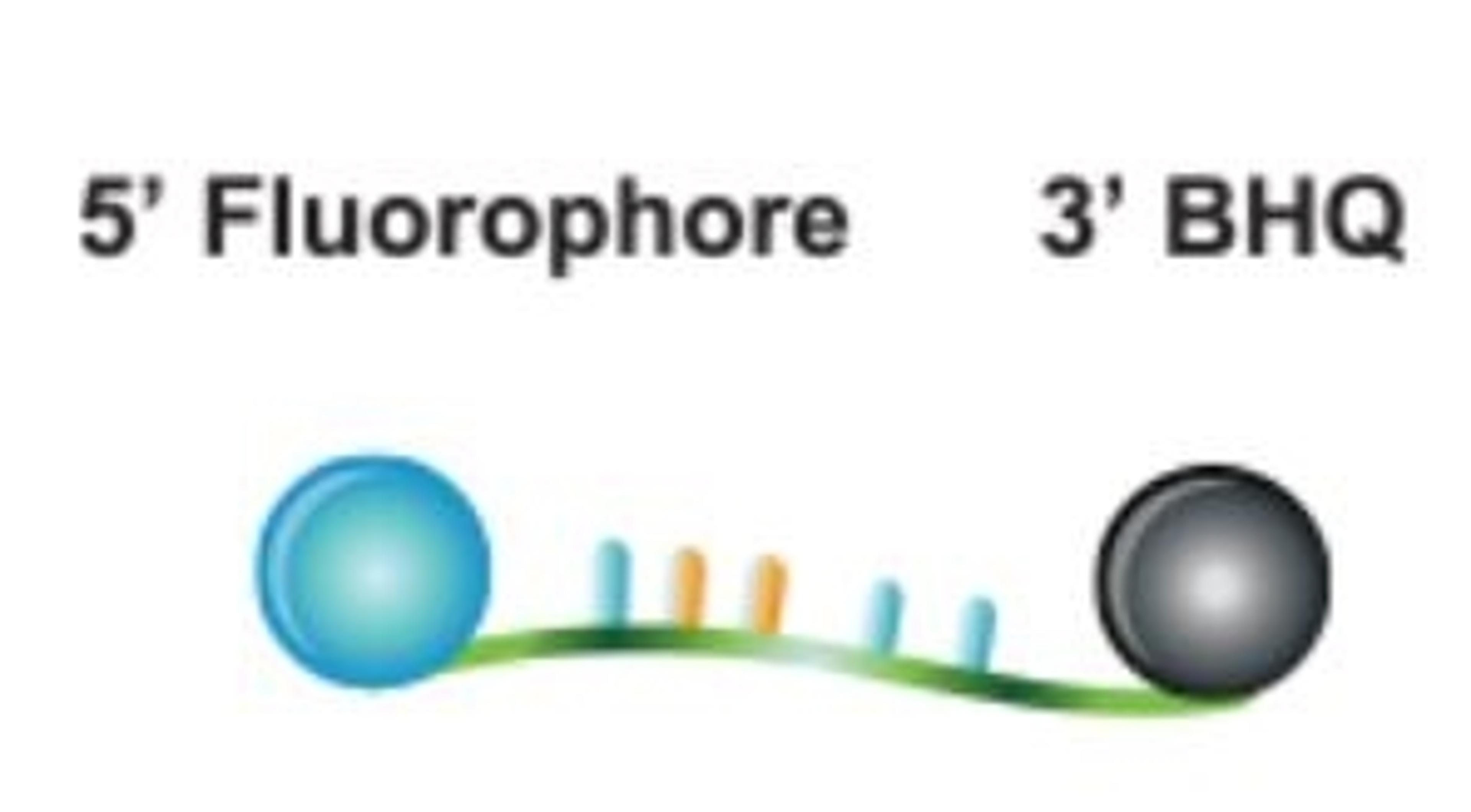
The DNA probing element incorporates a self-complementary stem sequence at the 3' and 5' ends that holds the target specific region in a hairpin loop. A fluorophore molecule is covalently linked to the 5' end and, the dark quencher, a Black Hole Quencher dye, is linked to the 3' end of the loop structure. The fluorophore and BHQ dye are held in close physical proximity by the stem structure to ensure efficient quenching prior to thermal cycling.
Target-specific primer
The target-specific PCR primer is covalently linked to the 3' end of the DNA probing sequence through a blocking moiety. This moiety prevents polymerase extension into the probing element which could manifest as false positives
Top tips for robust qPCR results
Reproducibility issues are a common pain point experienced by scientists carrying out quantitative polymerase chain reaction (qPCR), or real-time PCR, with challenges being linked to suboptimal design, probe selection, or workflow execution. The inconsistencies impact not only data quality, but also wasted time, lost samples and delayed research milestones.
A successful qPCR experiment relies on a well‑designed workflow, from high‑quality sample preparation to optimized reagents, primers, and probes, supported by thoughtful assay design that avoids issues like nonspecific binding or inefficient amplification. Expert probe‑selection strategies enhance specificity, especially in multiplexing, while effective troubleshooting helps diagnose common problems such as poor efficiency, primer‑dimers, or high variability. By applying key optimization techniques, researchers can achieve sensitive, accurate, and reproducible qPCR results.
Download this SelectScience guide to explore:
- How to overcome challenges in qPCR assay design
- A guide to selecting the right qPCR probe
- The best way to troubleshoot your qPCR
- Tips for qPCR optimization and validation
Resource details:
- Document type: SelectScience guide
- Page count: 49
- Read time: 69 mins
- Edition: 1st
7 innovative qPCR probe types explained
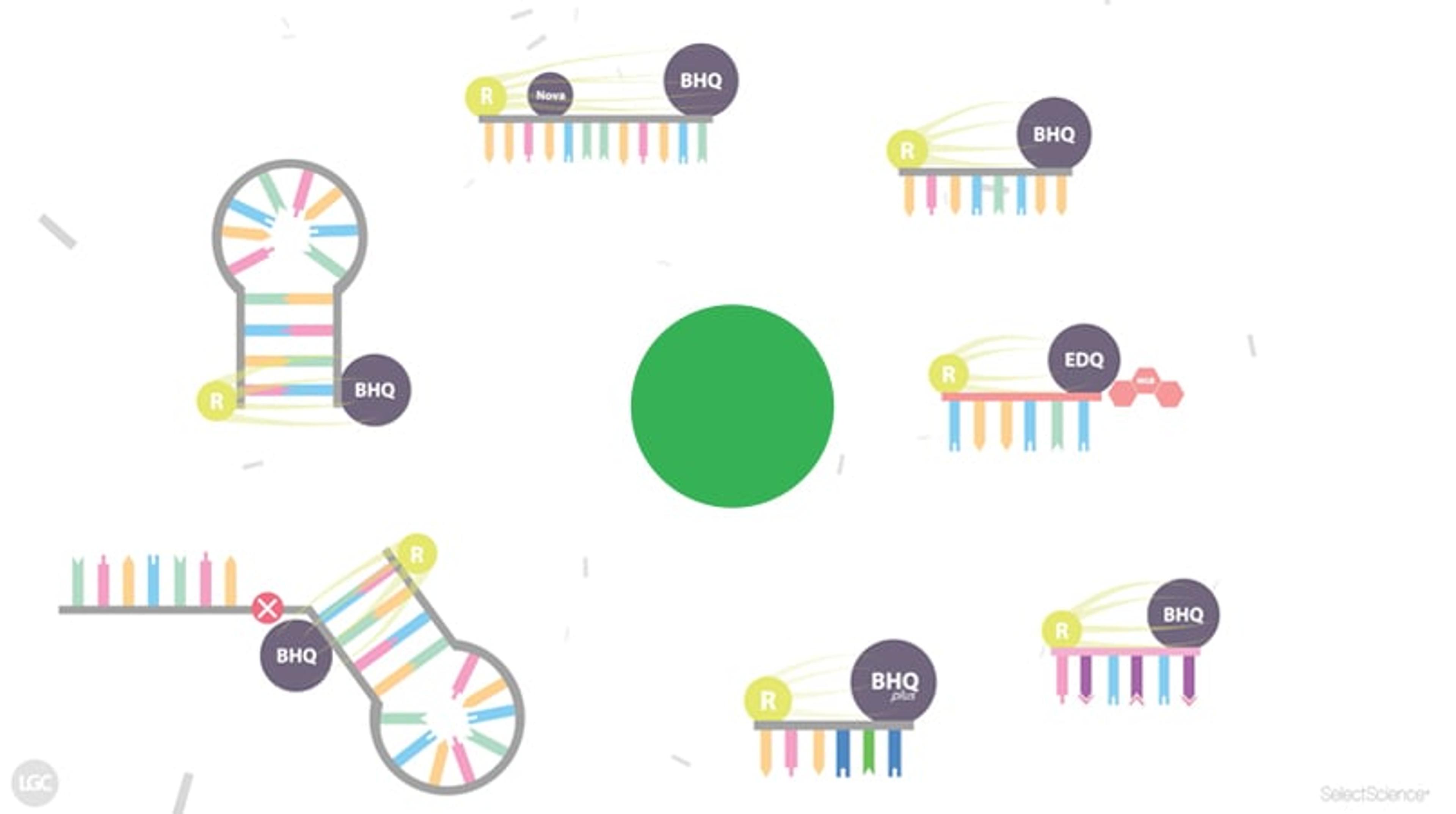
This comprehensive walkthrough animation outlines the latest developments in probe technologies in the qPCR field, highlighting the mechanisms of seven different real-time PCR probe and primer formats:
- Dual-labelled hydrolysis probes [00:19]
- BHQnova™ Probes [01:17]
- Minor groove binder (MGB) probes [01:49]
- Locked nucleic acid (LNA) probes [02:31]
- BHQplus™ Probes [03:15]
- Molecular Beacons [04:45]
- Scorpions™ Primers [06:00]