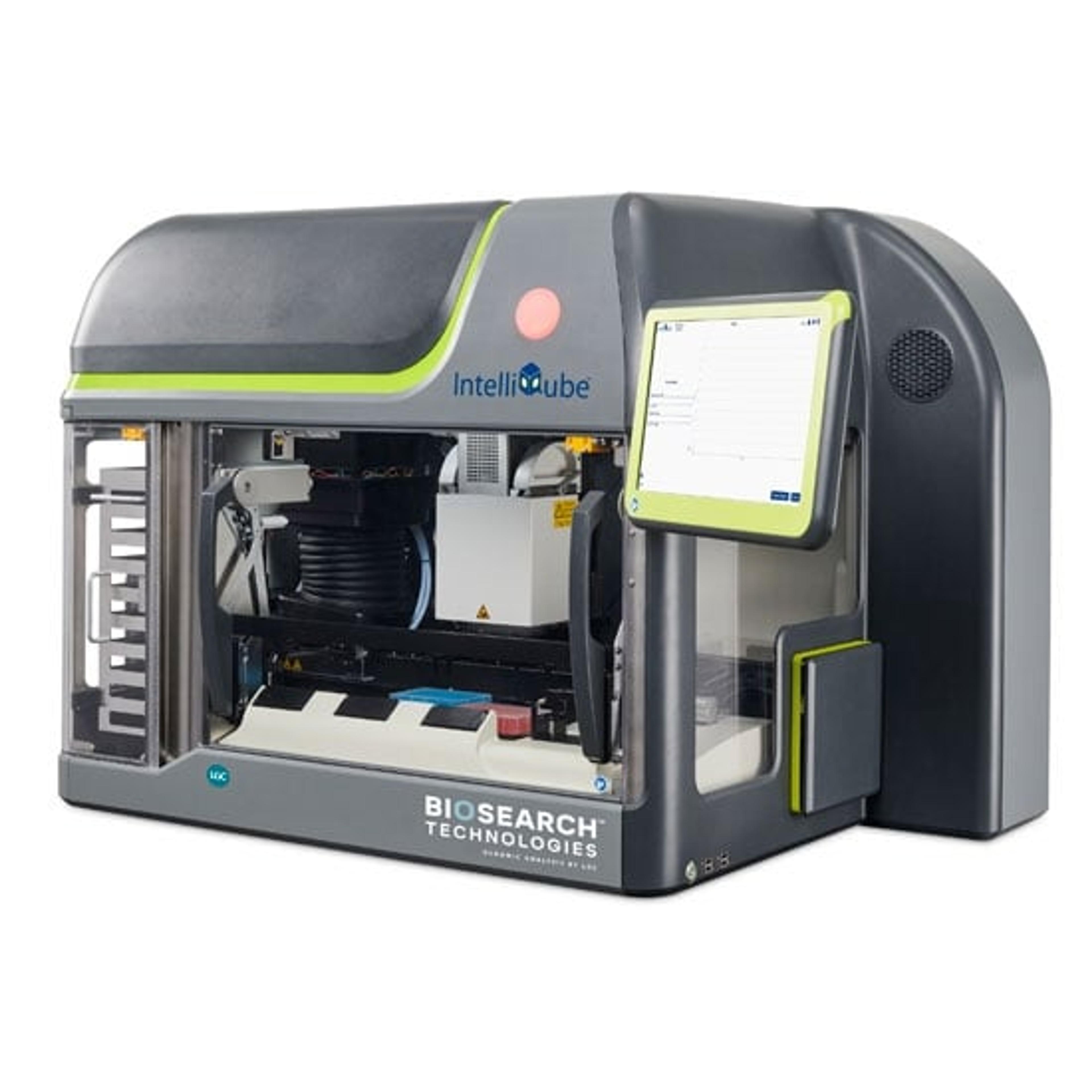
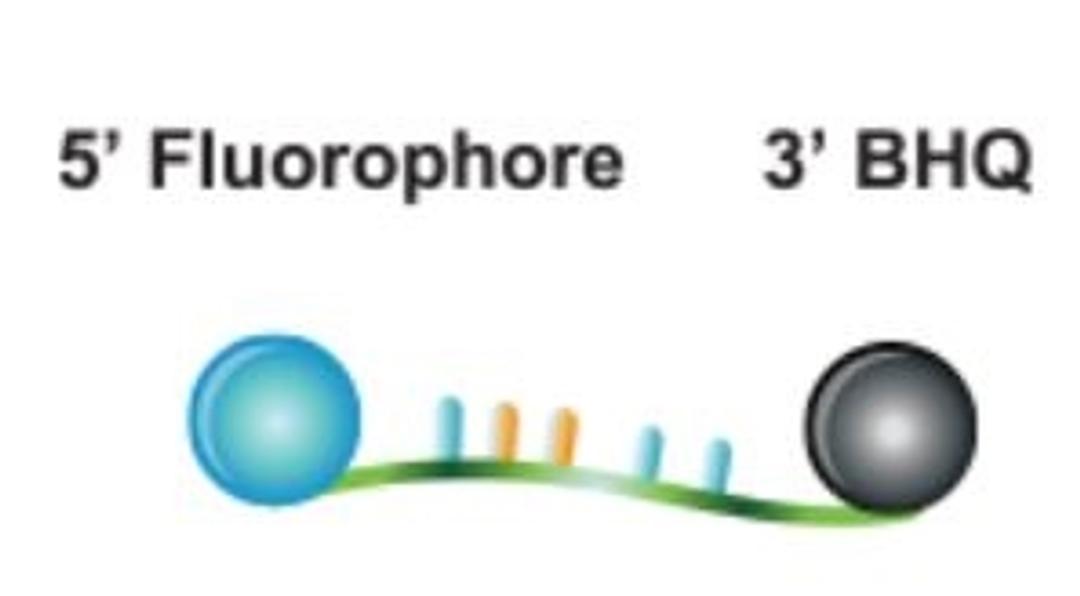
BHQplus™ Probes
Compact PCR probes for detecting difficult target sequences
Easy to design and use and the best value out there!
diagnostic testing
The probes are easy to use and the best value on the market. You can scale up the production size by contacting LGC and they are more than happy to synthesize any size you need and have it delivered in a timely fashion. I would like to see the BHQplus PCR probes come is a larger selection of fluorophores similar to their dual-labeled probes. In multiplexes we so see some spectral overlap between the FAM and Cal Fluor orange even with spectral calibration of the instrument which when you are looking for an absence of signal is very difficult to interpret.
Review Date: 1 Aug 2017 | LGC Biosearch Technologies
Great Results
Viral Load
Easy to use, good QC and fast rate for production.
Review Date: 31 Jul 2017 | LGC Biosearch Technologies
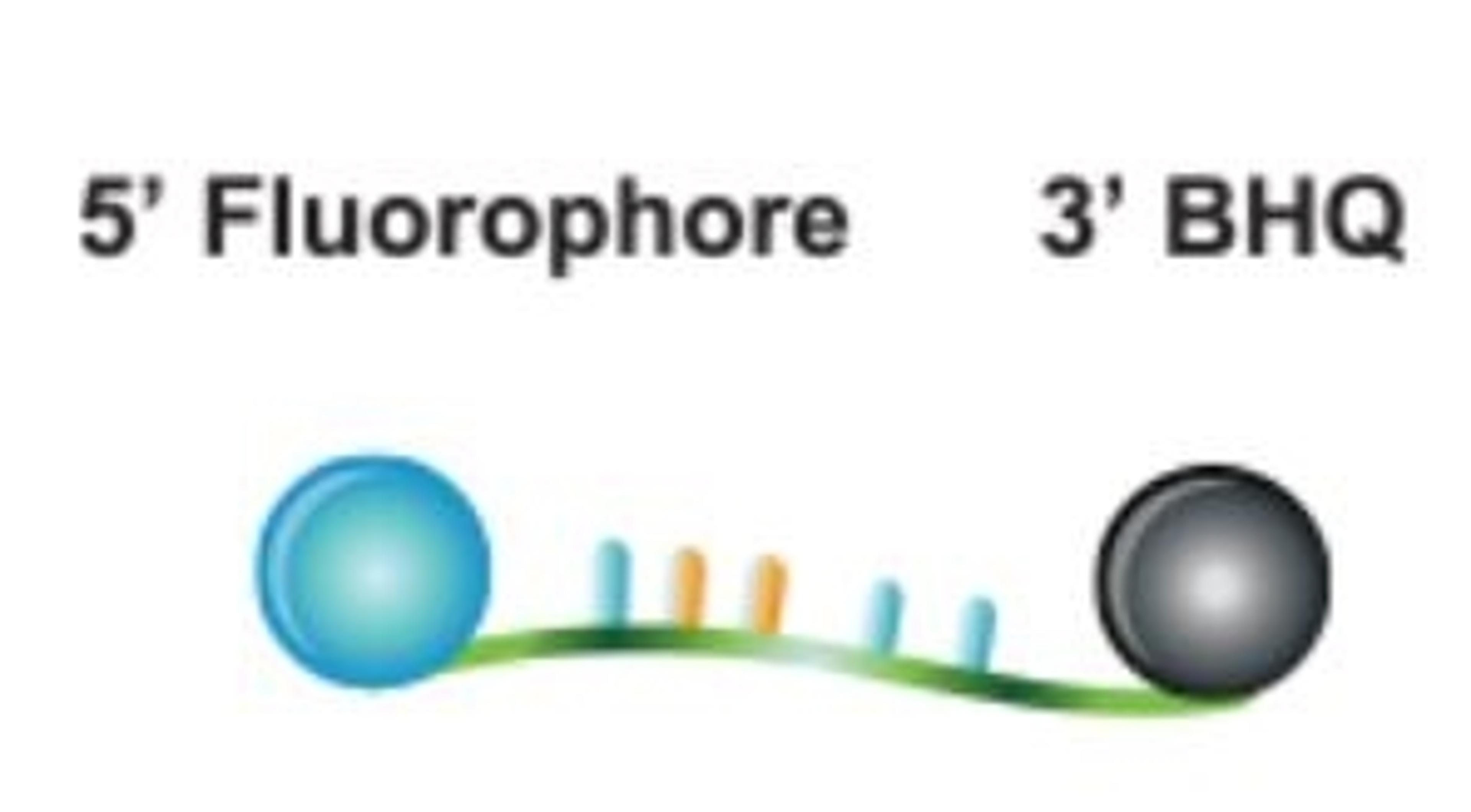
The BHQplus™ probe is a novel compact probe for qPCR, modified with a duplex stabilizing technology and terminated with Biosearch’s Black Hole Quencher™ dye. What differentiates the BHQplus probe from other traditional probes is that it permits the design of shorter oligonucleotides, typically 15 - 25 bases in length, for detecting more difficult target sequences.
The enhanced specificity of BHQplus probes make them perfect for genotyping single nucleotide polymorphisms.
Features:
- Small but powerful: duplex stabilizers permit the design of shorter dual-labeled probes
- High fidelity: shorter sequences offer an increased barrier against hybridizing to a mismatch
- Proven performance: the Black Hole Quencher dye extinguishes fluorescence until target hybridization
- Bottom line: An economical alternative for probe-based SNP Genotyping
Top tips for robust qPCR results
Reproducibility issues are a common pain point experienced by scientists carrying out quantitative polymerase chain reaction (qPCR), or real-time PCR, with challenges being linked to suboptimal design, probe selection, or workflow execution. The inconsistencies impact not only data quality, but also wasted time, lost samples and delayed research milestones.
A successful qPCR experiment relies on a well‑designed workflow, from high‑quality sample preparation to optimized reagents, primers, and probes, supported by thoughtful assay design that avoids issues like nonspecific binding or inefficient amplification. Expert probe‑selection strategies enhance specificity, especially in multiplexing, while effective troubleshooting helps diagnose common problems such as poor efficiency, primer‑dimers, or high variability. By applying key optimization techniques, researchers can achieve sensitive, accurate, and reproducible qPCR results.
Download this SelectScience guide to explore:
- How to overcome challenges in qPCR assay design
- A guide to selecting the right qPCR probe
- The best way to troubleshoot your qPCR
- Tips for qPCR optimization and validation
Resource details:
- Document type: SelectScience guide
- Page count: 49
- Read time: 69 mins
- Edition: 1st
7 innovative qPCR probe types explained
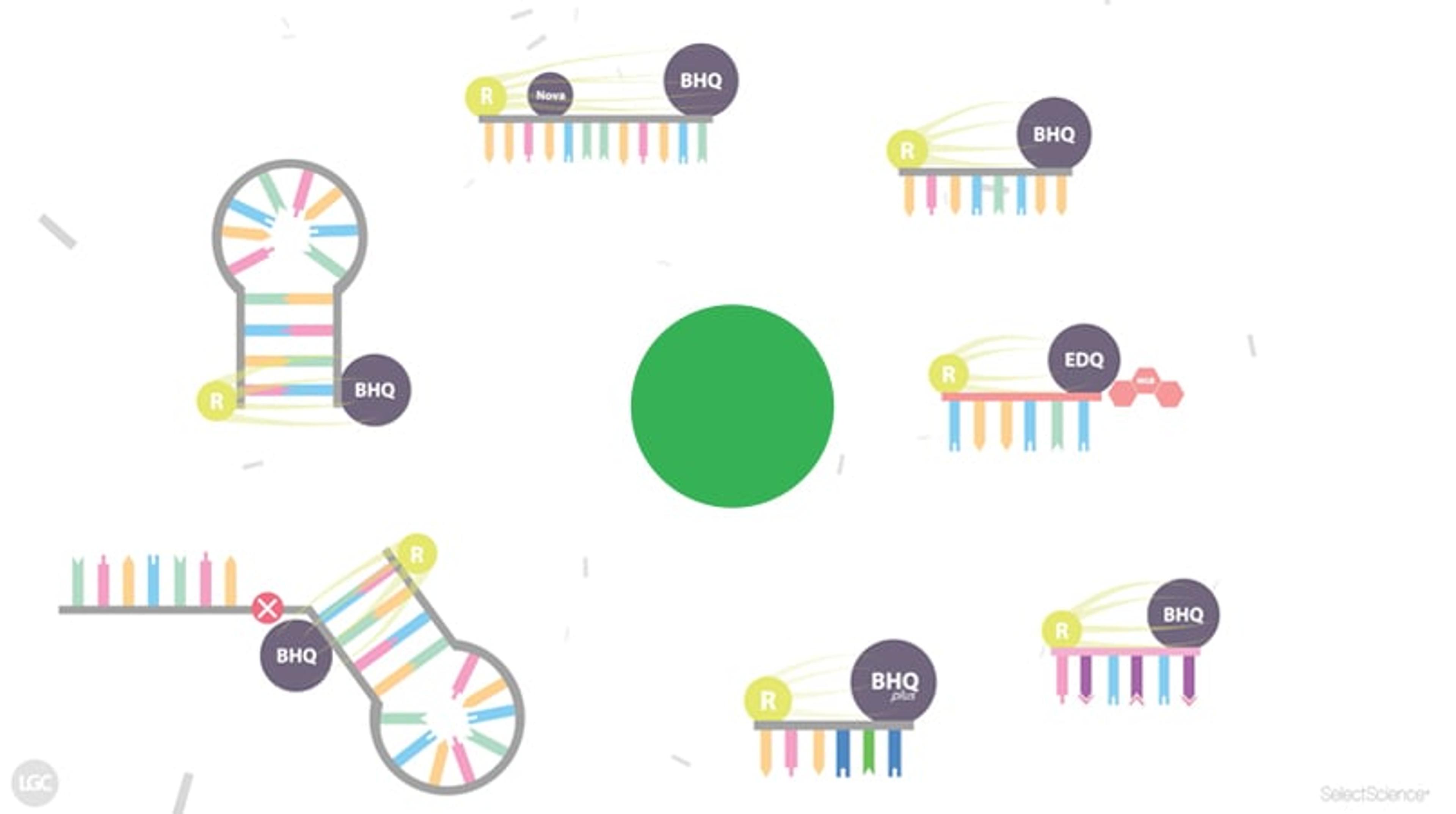
This comprehensive walkthrough animation outlines the latest developments in probe technologies in the qPCR field, highlighting the mechanisms of seven different real-time PCR probe and primer formats:
- Dual-labelled hydrolysis probes [00:19]
- BHQnova™ Probes [01:17]
- Minor groove binder (MGB) probes [01:49]
- Locked nucleic acid (LNA) probes [02:31]
- BHQplus™ Probes [03:15]
- Molecular Beacons [04:45]
- Scorpions™ Primers [06:00]
Combating antibiotic-resistant sexually transmitted infections with molecular diagnostics
Explore how a partnership between a lead researcher in sexual health and a molecular diagnostics innovator helped to overcome the therapeutic hurdles that come with antibiotic resistance
Transition to molecular diagnostics: How agricultural expertise has helped to keep COVID-19 positive rates low
Making the most of agrigenomic expertise and existing technologies to enable a rapid response to the pandemic