KAPA Hyper Prep Kits
Versatile, streamlined solution for DNA library preparation for Illumina sequencing.

The supplier does not provide quotations for this product through SelectScience. You can search for similar products in our Product Directory.
The novel one-tube chemistry, optimally formulated and evolved enzymes enable higher yields of adapter-ligated library and lower amplification bias. This translates to higher library diversity, lower duplication rates and more uniform coverage, particularly for FFPE and low-input samples.
Improve library yields and sequence quality:
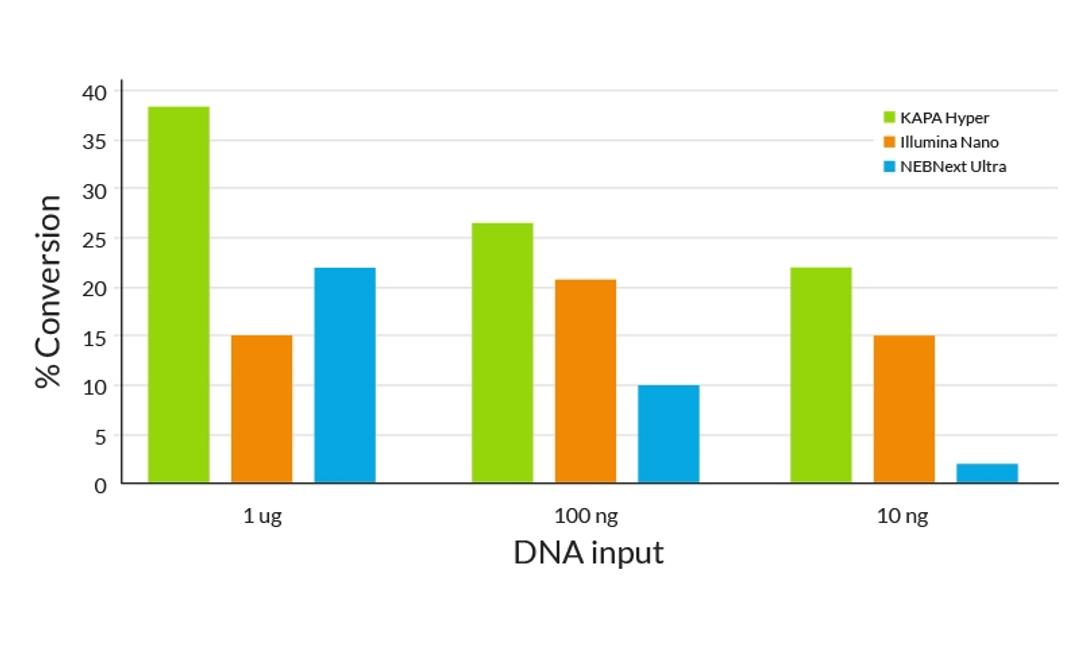
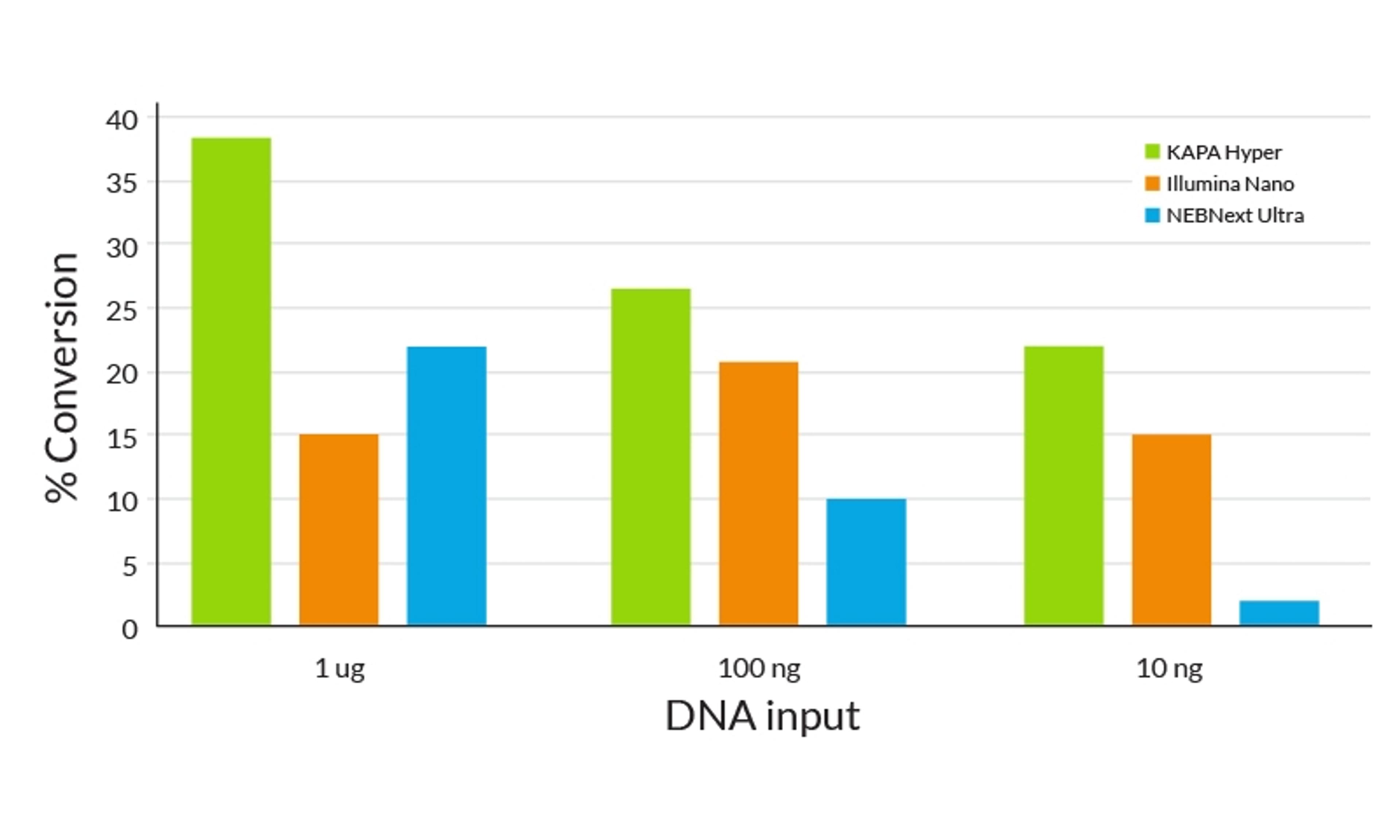
- Higher library yields translate to greater molecular complexity
- Fewer cycles of amplification with KAPA HiFi DNA Polymerase results in lower duplication rates and improved coverage
Create high-quality libraries from FFPE samples:
- Generate high-quality libraries from 250 ng FFPE DNA or less
- Significantly lower duplication rates and higher coverage
Complete library construction in less than 3 hours:
- One-tube workflow with minimal cleanup steps reduces overall time, and minimizes hands-on time
- Sample-to-library in <2 hours for PCR-free workflows, or <3 hours with amplification
- Fewer handling steps lead to improved consistency and reproducibility
Improve performance with low-input samples:
- Generate more sequence-quality library molecules without increasing adapter-dimers
- Increase yields without inducing size bias
Improving Data Quality and Turnaround Times for Targeted Next-Generation Sequencing of FFPE DNA using a KAPA HyperPlus/SeqCap EZ workflow
This application note evaluates the KAPA HyperPlus Kit from Kapa Biosystems for enzymatic DNA fragmentation and library construction. Formalin-fixed paraffin-embedded (FFPE) tissue is an important source of DNA for cancer genomic studies and clinical diagnostics. The KAPA HyperPlus Kit with integrated enzymatic fragmentation streamlines the SeqCapTM EZ Target Enrichment workflow, yielding high quality libraries.
Fast Turnaround Times and Improved Data Quality for Microbial Whole-Genome Sequencing with a Novel, Single-Tube Enzymatic Fragmentation
Next-generation whole genome sequencing of microbes demands rapid, robust, and scalable library construction workflows, capable of generating high-quality sequence data across a wide range of genome sizes, complexities and genomic GC content. This application note describes a streamlined library preparation method that results in minimal bias, high uniform coverage, and facilitates de novo assembly of microbial genomes.
A Streamlined Solution for the Construction of ChIP-Seq Libraries from Picogram Amounts of DNA
ChIP-Seq, chromatin immunoprecipitation followed by next-generation sequencing (NGS), is a valuable method enabling the study of DNA interactions with proteins, such as transcription factors and histone modifications. In ChIP-Seq workflows, proteins bound to DNA fragments of interest are enriched through formaldehyde crosslinking and targeted antibody selection, commonly known as immunoprecipitation. The enriched DNA is then purified and used as input into NGS library construction in preparation for sequencing. This application note demonstrates the use of the KAPA Hyper Prep Kit to offer a streamlined and effective solution for the construction of ChIP-Seq NGS libraries from ultra-low DNA inputs.
KAPA Hyper Prep and HiFi Uracil+ Kits Allow for Reduced Representation Bisulfite Sequencing of Ultra-low Amounts of DNA in B Cells
This application note provides a robust methodology for RRBS analysis using the streamlined chemistry of the KAPA Hyper Prep Kit and HiFi Uracil+ HotStart DNA Polymerase, and demonstrates its utility in accurately quantifying the DNA methylation status from sample-limiting conditions. Methyl-Seq is a powerful tool for understanding genome-wide methylation with base-pair resolution. Several strategies have been developed to understand the methylation status of a genome, including reduced representation bisulfite sequencing (RRBS). Bisulfite treatment results in a significant decrease in DNA input and quality; therefore, the construction of high-quality libraries and efficient downstream amplification is critical.
Application of NGS in the Diagnostic Lab on Archival FFPE Tissues
Hosted by Kapa Biosystems at AMP2015, Dr. Brian Walker of the Royal Marsden Hospital presents at a work shop entitled "Application of NGS in the diagnostic lab on archival FFPE tissues".
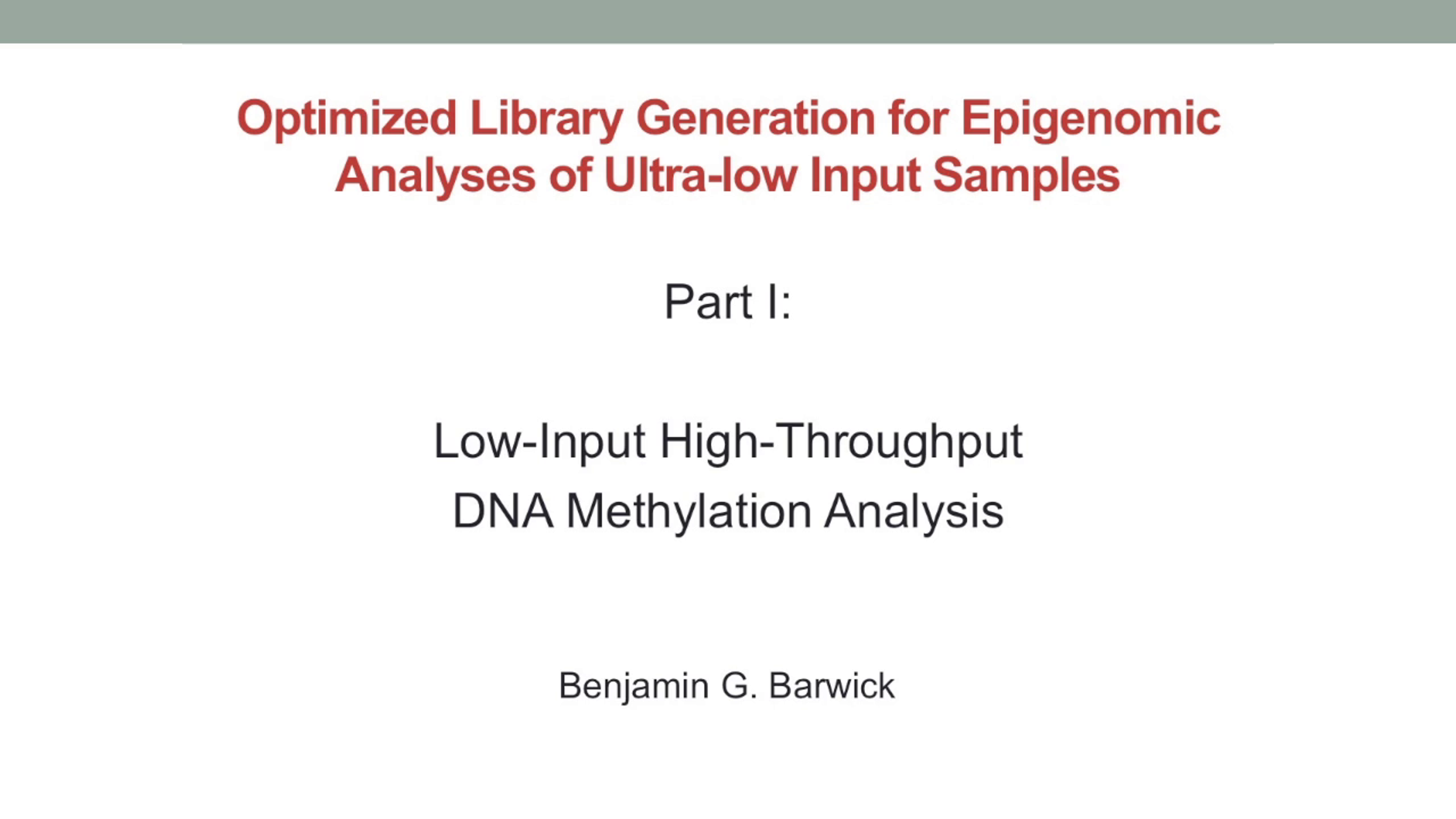
Low-Input DNA Methylation Analysis at Emory University
DNA methylation is essential for mammalian development and is often dysregulated in disease. Understanding the dynamics of DNA methylation can thus provide insight into disease etiology and serve as a biomarker. Base-pair resolution analysis of DNA methylation is facilitated by sequencing of bisulfite treated DNA, but this process can be laborious and expensive. Reduced representation bisulfite sequencing (RRBS) uses restriction enzymes to enrich parts of the genome where CpG dinucleotides occur, thus decreasing sequencing costs. In this video presentation from ASHG 2015, Ben Berwick of Emory University presents a robust methodology for RRBS analysis using methods from Kapa Biosystems.