duet multiomics solution evoC
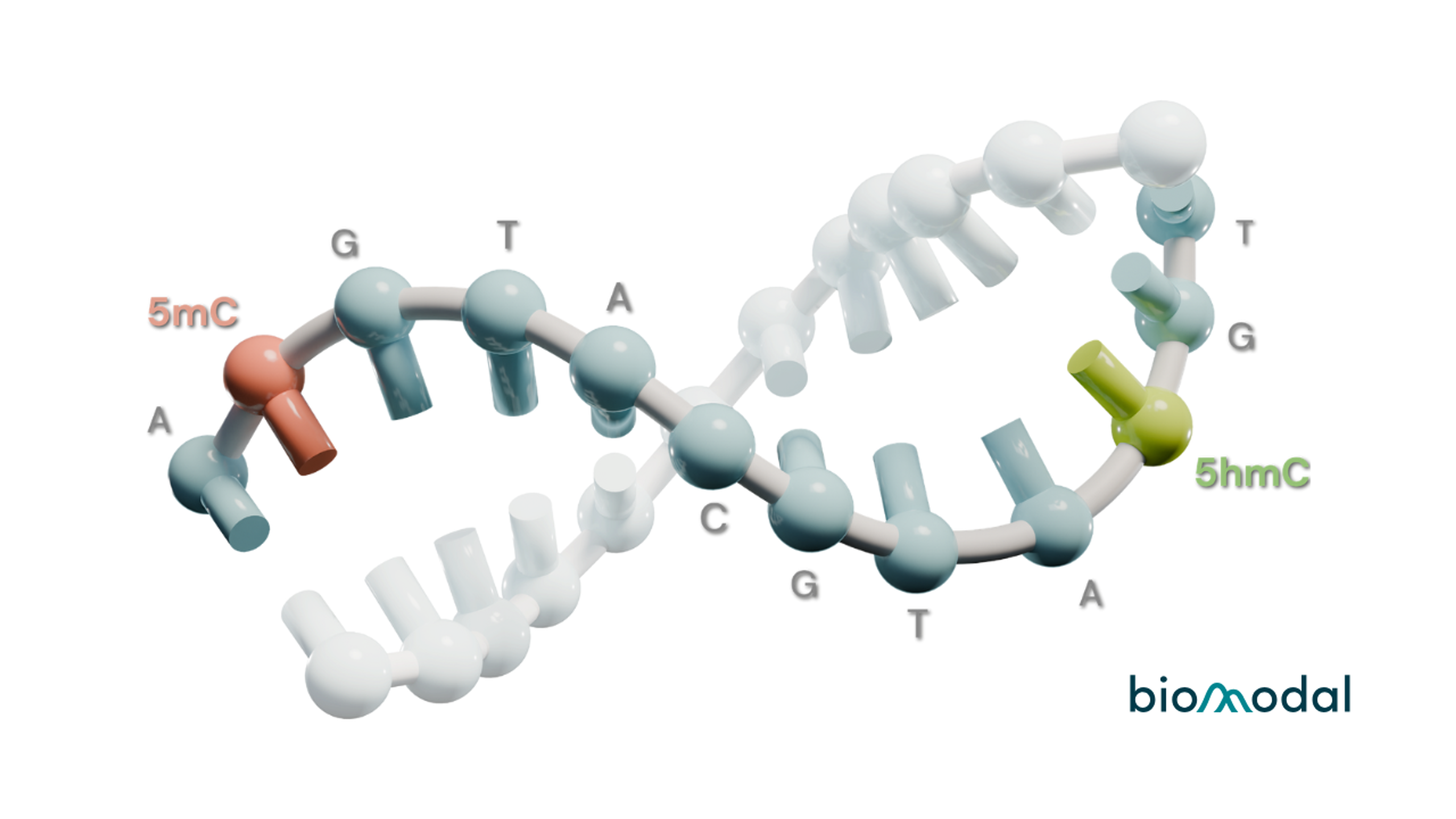
Your single solution for multiomics data - Uncover a true 6-base genome for unprecedented insights into current and future biological states. Access all four canonical bases and distinguish 5‑methylcytosine (5mC) and 5‑hydroxymethylcytosine (5hmC) from a single 5ng DNA sample, in a single experiment, with high accuracy.

The supplier does not provide quotations for this product through SelectScience. You can search for similar products in our Product Directory.
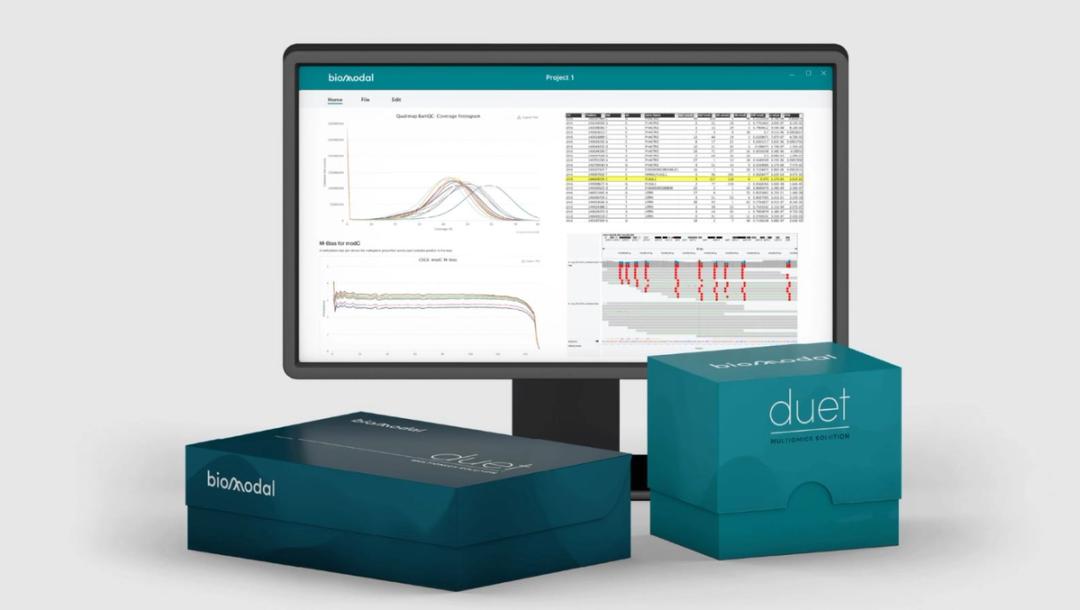
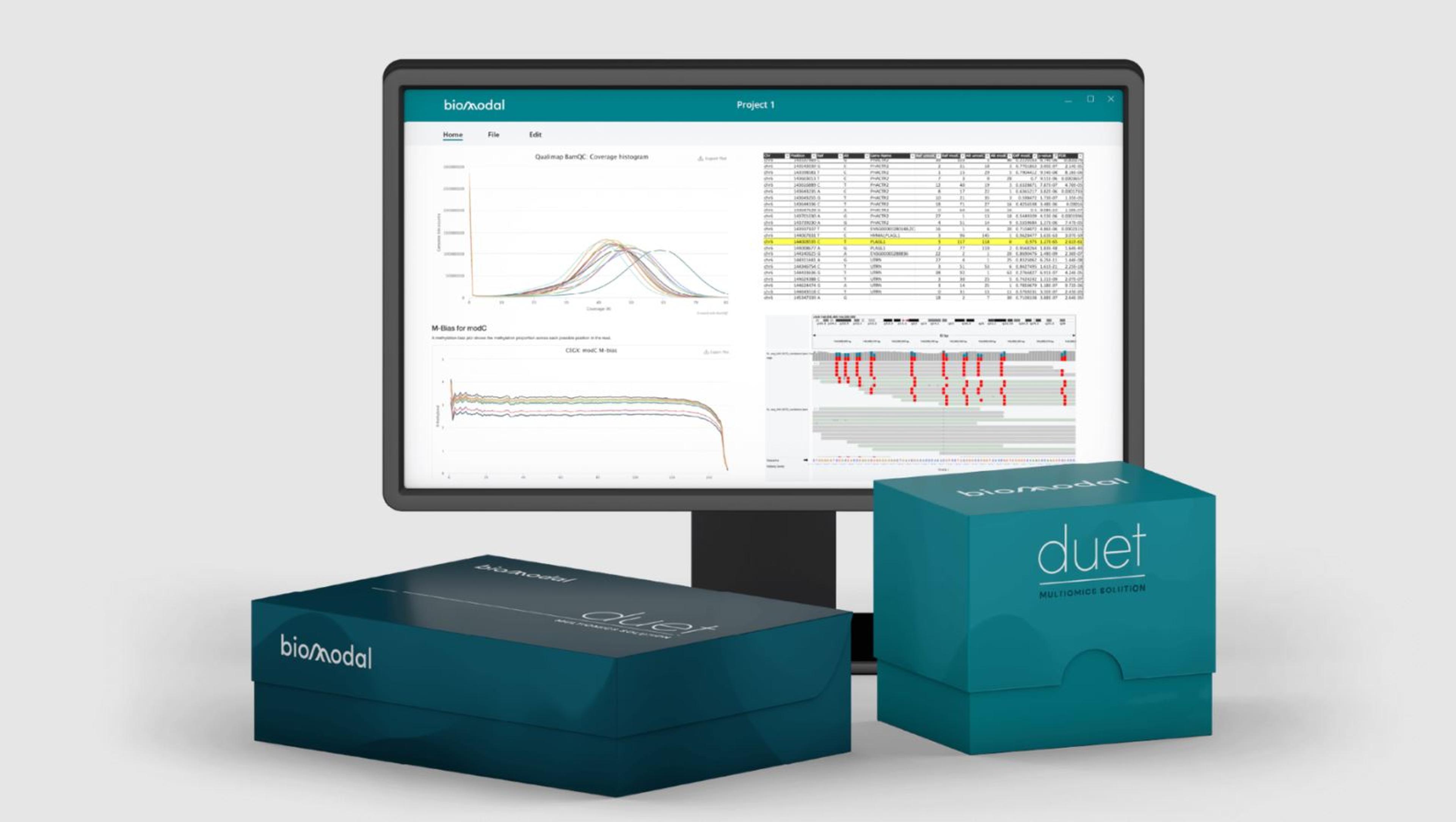
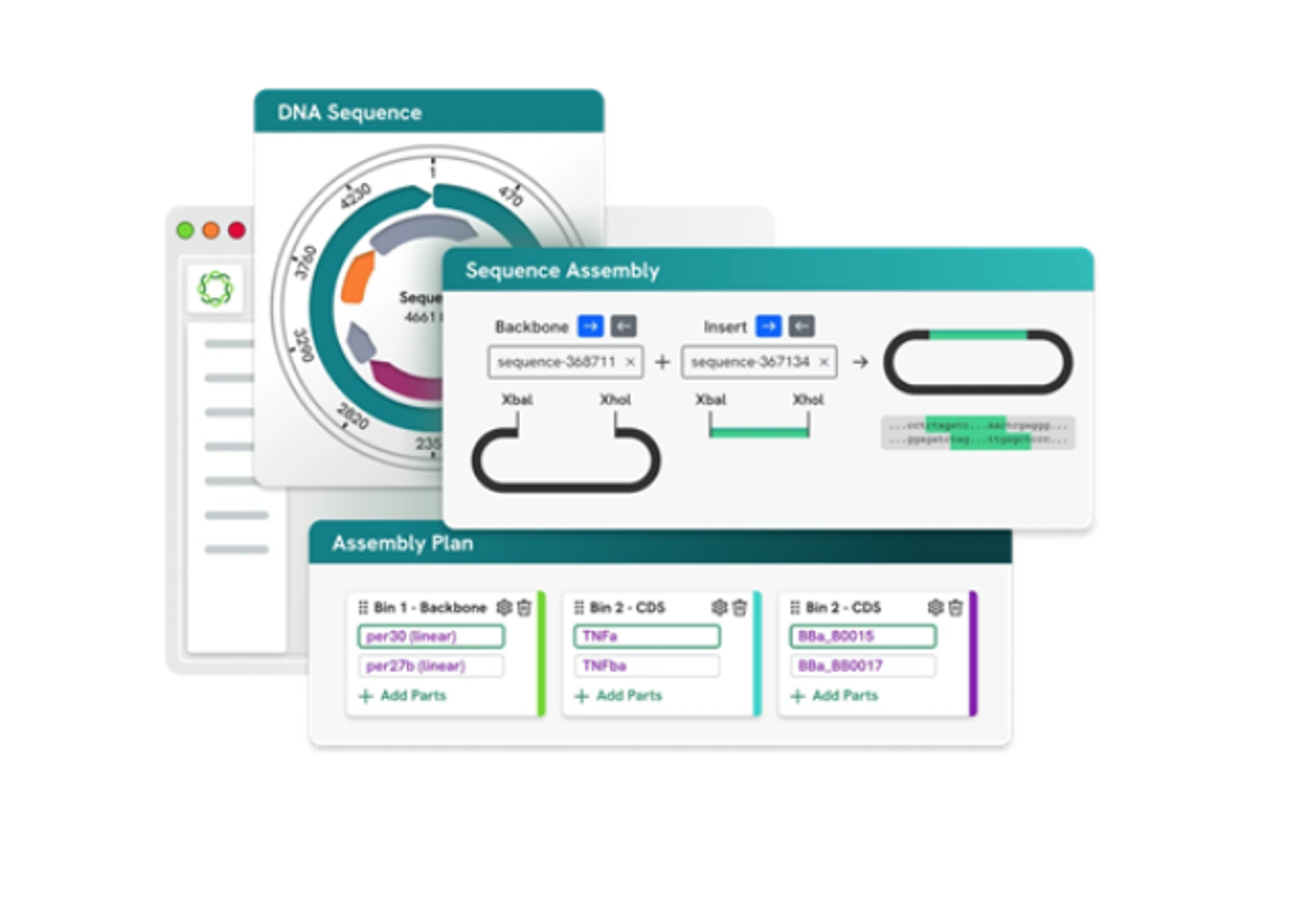
duet multiomics solution evoC is comprised of a pre-sequencing workflow requiring as little as 10ng gDNA or 5ng cfDNA that seamlessly integrates with NGS sequencing platforms and a post-sequencing software package to accelerate data analysis. The post-sequencing software trims, aligns, and annotates to generate analysis-ready BAM, VCF, BedMethyl, and ASM files. Additional plugins include proprietary analysis software that facilitates analysis of multimodal data at scale.
The 6-base genome provides a combined genome and methylome with full discrimination of 5mC and 5hmC for genetic sequence analysis, variant identification, and DNA methylation analysis.
The 6-base genome data delivers:
- Somatic vs. germline mutations
- Variant associated methylation
- Fragment profiles
Methylation status of control elements - Differentially methylated loci and regions
Mapping human tissues with 6-base sequencing
5-methylcytosine (5mC) and 5-hydroxymethylcytosine (5hmC), often referred to as the fifth and sixth DNA bases, exhibit distinct tissue-specific distributions and can serve as molecular fingerprints. Using 6-base sequencing with duet evoC, a multiomic platform capable of simultaneously detecting all four canonical DNA bases alongside 5mC and 5hmC, researchers can comprehensively profile the 6-base genome across multiple tissues.
Data analysis with Biomodal’s XPLR software reveals biologically relevant structures within the methylation landscape. Differential methylation region (DMR) analysis further identifies tissue-specific 5mC and 5hmC patterns linked to key pathways and regulatory programs, providing deeper insight into epigenetic regulation and tissue identity.
Large diagnostics laboratory adopts 6-base sequencing to improve disease characterisation
The Munich Leukemia Laboratory (MLL) adopted biomodal’s duet evoC 6-base sequencing technology to enhance disease characterization through high-resolution methylation profiling. The technology demonstrated improved ease of use, deep genomic coverage, and low background noise, providing promising results for patient stratification and biomarker discovery in leukemia diagnostics.
Measuring genetics, 5-mC and 5-hmC at single-base resolution
Explore he duet multiomics solution evoC, an innovative sequencing technology that simultaneously resolves the four genetic bases and distinguishes between cytosine modifications 5-methylcytosine (5mC) and 5-hydroxymethylcytosine (5hmC) (six-base calling). By integrating genetic and epigenetic data at high accuracy, evoC enables deeper insights into gene expression, chromatin accessibility, and enhancer states. Using mouse embryonic stem cells, this method reveals the dynamic interplay between DNA modifications and transcriptional regulation.
Reveal the power of the 6-base genome from a single DNA molecule
Discover how 6-base genome analysis, integrating 5-methylcytosine (5mC) and 5-hydroxymethylcytosine (5hmC) insights, revolutionizes genomic research. Gain a comprehensive view of genetic and epigenetic interactions to predict disease progression, identify biomarkers, and guide clinical decisions for cancer, neurodegenerative diseases, and precision medicine.
Multiomic 6-base data from cell-free DNA enhances the performance of liquid biopsy classifiers
Early cancer detection has the potential to significantly improve treatment outcomes and survival rates. Investigate the roles of 5-methylcytosine (5mC) and 5-hydroxymethylcytosine (5hmC) as biomarkers for early-stage colorectal cancer (CRC) detection in cell-free DNA (cfDNA), using duet evoC for whole genome 6-base sequencing. Discover how the results support the hypothesis that distinguishing between 5mC and 5hmC can improve the sensitivity of liquid biopsy tests for early cancer detection.
Comprehensive, simultaneous genetic and epigenetic profiling of cell-free DNA in hepatocellular carcinoma
Discover a cutting-edge study on hepatocellular carcinoma (HCC) using duet evoC multiomics, which provides simultaneous genetic and epigenetic profiling of cell-free DNA (cfDNA) in a single assay. This research explores copy number alterations (CNAs), SNPs, and DNA methylation patterns to enhance early cancer detection and precision medicine.
Genetic and epigenetic study of formalin-damaged DNA with duet multiomics solution evoC 6-base sequencing
Explore the impact of formalin fixation on DNA methylation profiles using the innovative 6-base sequencing technology, duet multiomics solution evoC. By analyzing formalin-compromised, FFPE, and fresh-frozen cancer samples, the research demonstrates that despite minor predictable changes, the method provides reliable genetic and epigenetic insights, preserving the biological integrity of stored samples.
Epigenetic inhibitors with PARPis: A novel therapeutic strategy for BRCA-proficient pancreatic cancer
Patient-derived pancreatic cancer organoids showed significant growth inhibition when treated with decitabine, an epigenetic inhibitor. Noticeably, the combination of decitabine with a PARP inhibitor showed pronounced synergism in inhibiting tumour organoid growth.
By employing the 6-base genome via duet evoC, researchers at the University of Maryland have identified differentially methylated genes that may be responsible for therapy resistance and tumor aggression in pancreatic cancer.
Epigenetic analysis of 6-base genome in Parkinson’s
The emergence of the ‘6-base genome’, which includes the modified bases 5-methylcytosine (5mC) and its oxidised form, 5-hydroxymethylcytosine (5hmC), marks a significant evolution in our understanding of genetic and epigenetic regulation.
In this webinar, Burleen Chhatwal from University College London, UK, demonstrates how biomodal’s 6-base genome supported the successful analysis of both 5mC and 5hmC modifications in Parkinson’s disease compared to controls in a pilot study.
The 6-base genome revolution
Dr. Chia-Lin Wei, Professor of Genome Sciences at University of Washington, shares how to unlock DNA methylation insights for neuroscience and cancer research