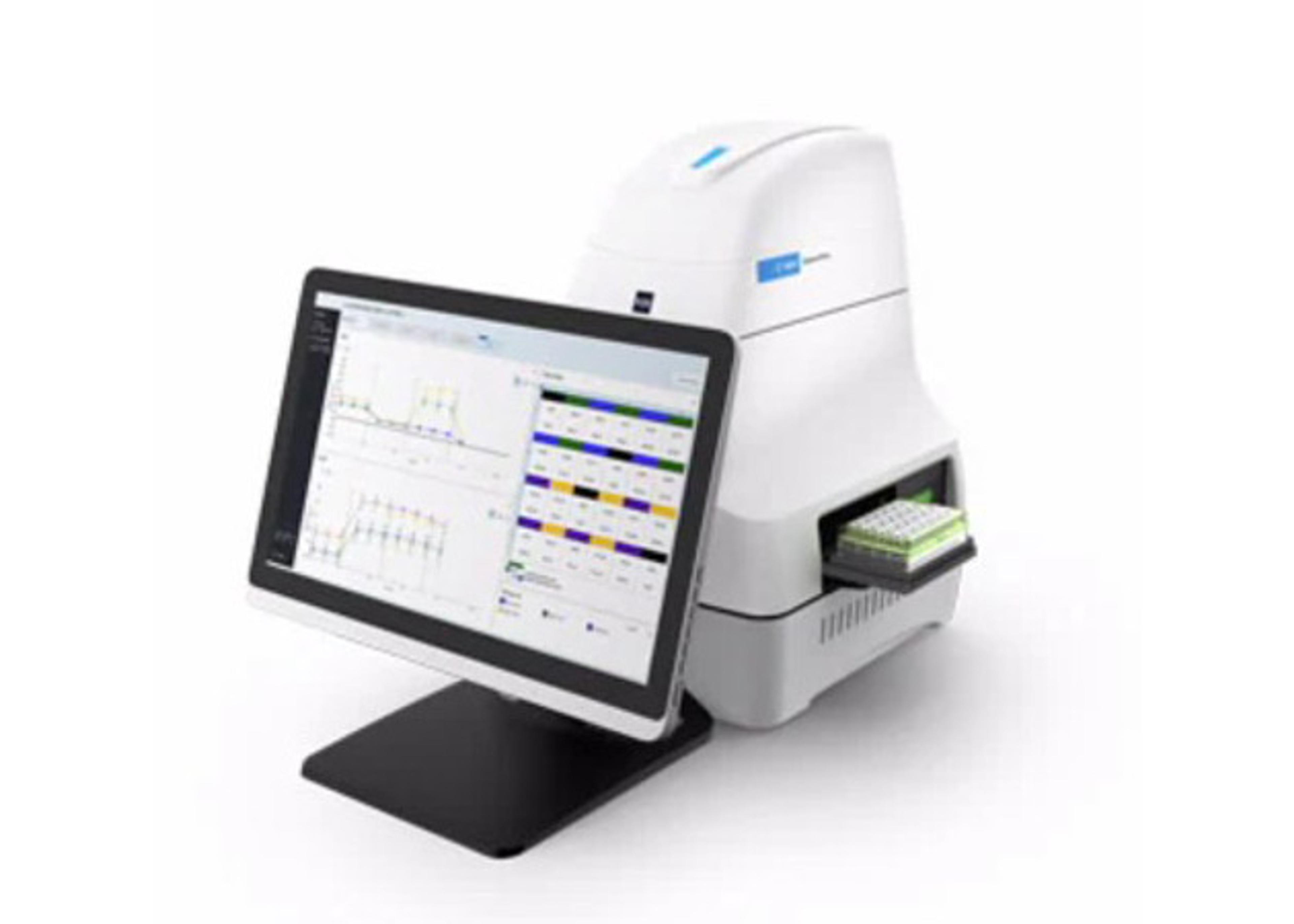
SureSelect Target Enrichment System Kit
Agilent's SureSelect market leading platform provides a complete portfolio of catalog and custom products, providing the flexibility you need, for your discovery to follow-up studies. Agilent™s SureSelect Target Enrichment System is a highly efficient hybrid selection technique for optimizing next-generation sequencing. It is available for the Illumina GA and HiSeq, Applied Biosystems SOLiD, and 454 Life Sciences FLX and GS J…
Highly recommended NGS library prep kit.
Targeted NGS
The SureSelect QXT Target Enrichment Kit is doing great and it is a really effective library prep kit for NGS. I used to get very high yield with this kit. I highly recommend it to anyone planning to do targeted NGS experiments. Also SureDesign tool is easy to use to design customized probe sets.
Review Date: 22 Jul 2019 | Agilent Technologies
Agilent's SureSelect market leading platform provides a complete portfolio of catalog and custom products, providing the flexibility you need, for your discovery to follow-up studies.
Agilent™s SureSelect Target Enrichment System is a highly efficient hybrid selection technique for optimizing next-generation sequencing. It is available for the Illumina GA and HiSeq, Applied Biosystems SOLiD, and 454 Life Sciences FLX and GS Junior NGS systems, including paired or single-end protocols.
SureSelect Target Enrichment System Kit Features:
- Custom design your kit through eArray.
- Highly automatable, easy design process.
- Kit sizes can be chosen to accommodate very small projects, up to thousands of samples.
- Contents include all reagents and customer-specified libraries.
Targeted Exome and Gene Panel Analysis from Low Input and FFPE DNA using Hybridization Capture for Cancer Genome Studies
Hybridization-based target enrichment techniques coupled with Next-Generation Sequencing (NGS) provide a useful and cost-efficient means to study disease specific target regions including whole exomes and gene panels. Clinical samples may be limited in input and of compromised quality due to formalin fixation, however most current NGS library preparation methods require 50-100 ng high quality DNA for such studies. This application note presents an efficient method, which enables high quality target enrichment and variant calling from inputs as low as 1-25 ng.
How automation and integrated sample QC enhance a targeted long-read-sequencing workflow
Tuesday, November 4, at 16:00 GMT | 17:00 CET | 11:00 EST | 8:00 PST
As the demand for greater accuracy and efficiency in genetic analysis grows, new approaches are needed to overcome the blind spots of conventional sequencing methods. This webinar will showcase a fully automatable workflow that integrates Agilent SureSelect hybrid capture, Agilent automated electrophoresis systems for sample quality control with PacBio HiFi long-read sequencing.
In addition to discussing workflow, we’ll share case studies where long reads play a fundamental role in identifying genetic variants that may be missed by conventional sequencing in research studies.
Join us to learn how targeted long-read sequencing:
- Focuses on genes and regions of interest in genetic research, while reducing costs and boosting sequencing depth at key loci.
- Overcomes the limitations of short-read technologies, including challenges with pseudogenes, homologous loci, structural variants, complex transcript isoforms, splicing events, and difficult regions such as repetitive or GC-rich sequences.
- Enhances both analytical performance and laboratory efficiency through automation.
Key learning objectives:
- Gain practical insights into implementing an automated targeted long-read sequencing workflow in the laboratory.
- Identify the genetic mutation types best suited for targeted long-read sequencing approaches.
- Understand the analytical advantages of long-read sequencing compared with conventional short-read methods.
Who should attend?
Lab directors, molecular pathologists, research scientists, clinical lab technicians, and anyone running or interested in running NGS workflows.
Certificate of attendance
If you attend the live webinar, you will automatically receive a certificate of attendance, including a learning outcomes summary, for continuing education purposes.
If you view the on-demand webinar, you can request a certificate of attendance by emailing editor@selectscience.net.
For Research Use Only. Not for use in diagnostic procedures. PR7003-294
Agilent's SureSelect DNA kit to accelerate capture-based enrichment library preparation
The new kit promises enhanced performance and flexibility to power cancer and genetics research