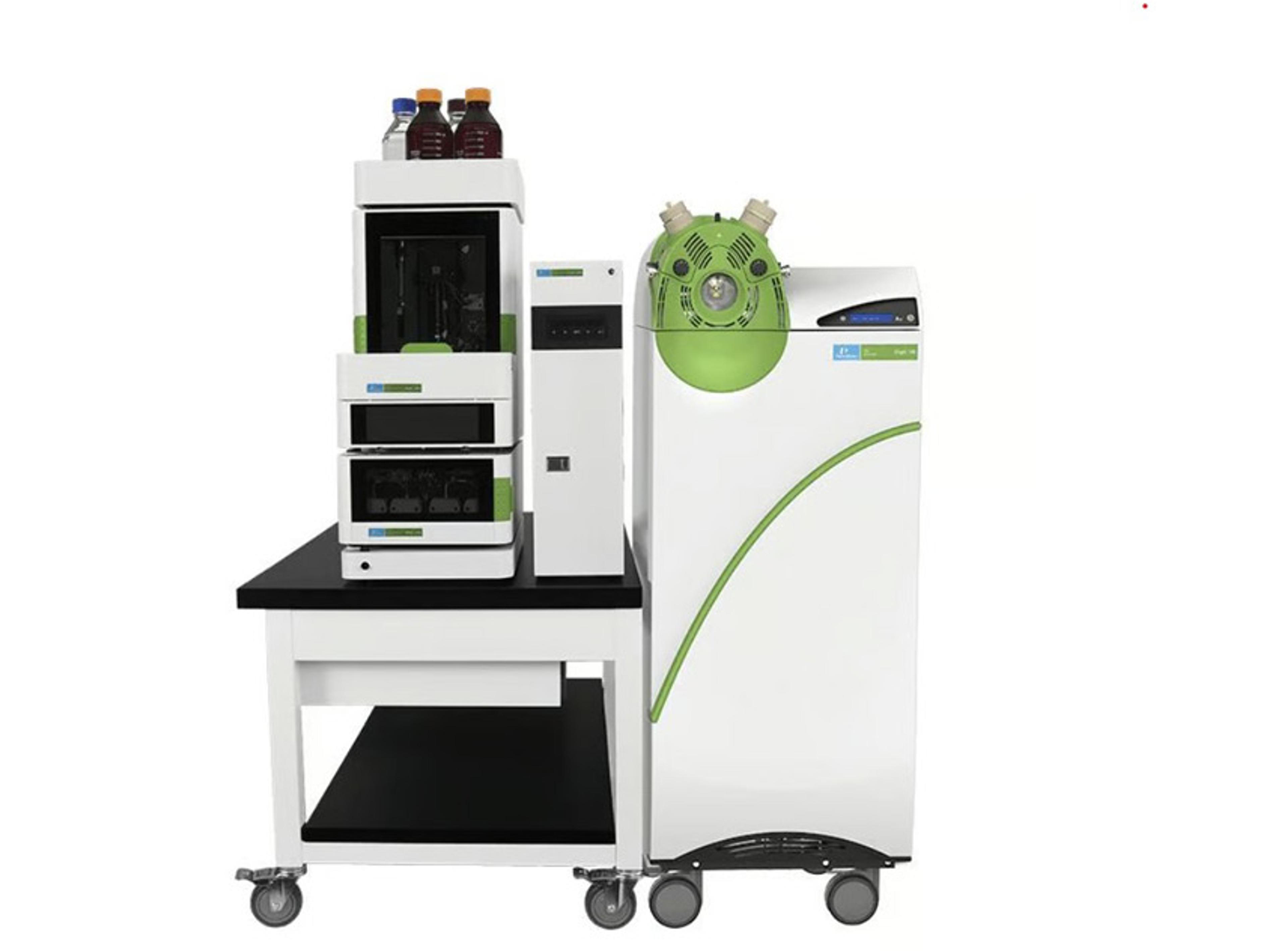
IdentityE High Definition Proteomics system
The Waters IdentityE High Definition Proteomics system defines the new standard for rigorous protein identification. The IdentityE High Definition Proteomics System is an UltraPerformance LC® (UPLC)/High-Bandwidth Mass Spectrometry (MS E) system solution suitable for confident protein identification designed to meet stringent requirements as outlined by leading scientific journals.The new Waters IdentityE High Definition Pro…

The supplier does not provide quotations for this product through SelectScience. You can search for similar products in our Product Directory.
The Waters IdentityE High Definition Proteomics system defines the new standard for rigorous protein identification.
The IdentityE High Definition Proteomics System is an UltraPerformance LC® (UPLC)/High-Bandwidth Mass Spectrometry (MS E) system solution suitable for confident protein identification designed to meet stringent requirements as outlined by leading scientific journals.
The new Waters IdentityE High Definition Proteomics System catalogs complex protein digest mixtures containing tens of thousands of peptides. The system features a Waters nanoACQUITY® UPLC System for high chromatographic peak capacity and retention time reproducibility, together with an electrospray tandem mass spectrometer - Waters Q-Tof Premier™ or Waters Synapt HDMS system - for high dynamic mass resolution and consistent over-sampling of complex protein digests – two hallmarks of high definition proteomics.
Waters comprehensive peptide ion accounting informatics mine deeper into UPLC/MS E data to identify more peptides/proteins with greater sequence coverage and statistical rigor than conventional LC/MS/MS methods.
This system solution is fully supported with quality control standards, defined methods, training and support to ensure reproducible operation and consistent results.
Determination of Quantitative Protein Signatures for Ductal Carcinoma (Breast Cancer) by LC/MS Proteome Analysis
In this study, a comprehensive MS-based discovery strategy is applied for a polygenic disease. The method employs the separation and detection of non-labelled tryptic fragments by means of an LC/MS acquisition. Instrumentation used included the Waters Expression High Definition Proteomics System, made up of the nanoACQUITY UPLC System and the Q-Tof Premier Mass Spectrometer.
Identification, Quanification and Annotation of Heart Membrane Proteins Using Label-Free Nanoscale LC /MS
Heart diseases resulting in heart failure are among the leading causes of morbidity and mortality in the Western world and can result from either systemic disease (e.g. hypertensive heart disease, ischaemic heart disease) or specific heart muscle disease (e.g. dilated cardiomyopathy (DCM)). Sub-proteome analysis of such disease sub-sets affords a reduction in sample complexity, potentially revealing biomarkers of cardiac failure that would otherwise remain undiscovered. Label-free nanoscale LC/MS is used in this study to validate a Triton X-114 based enrichment method for cardiac membrane proteins.
7 Popular Cancer Research Applications
Discover seven applications to improve your cancer research methods