PACBIO RS II
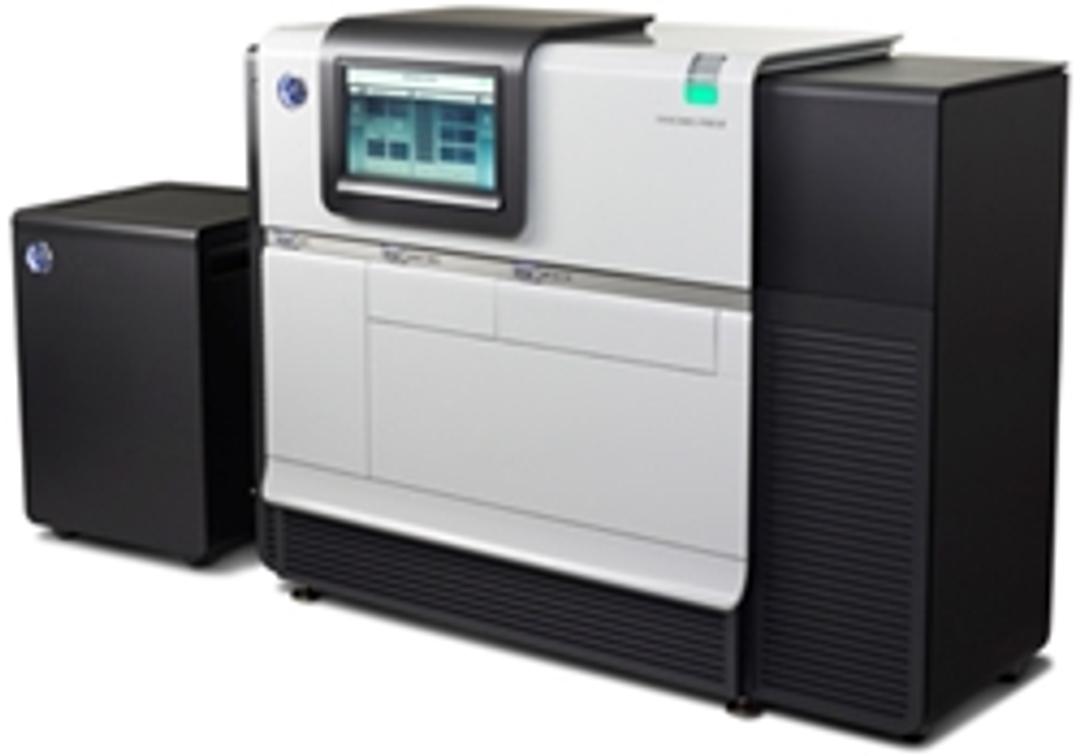
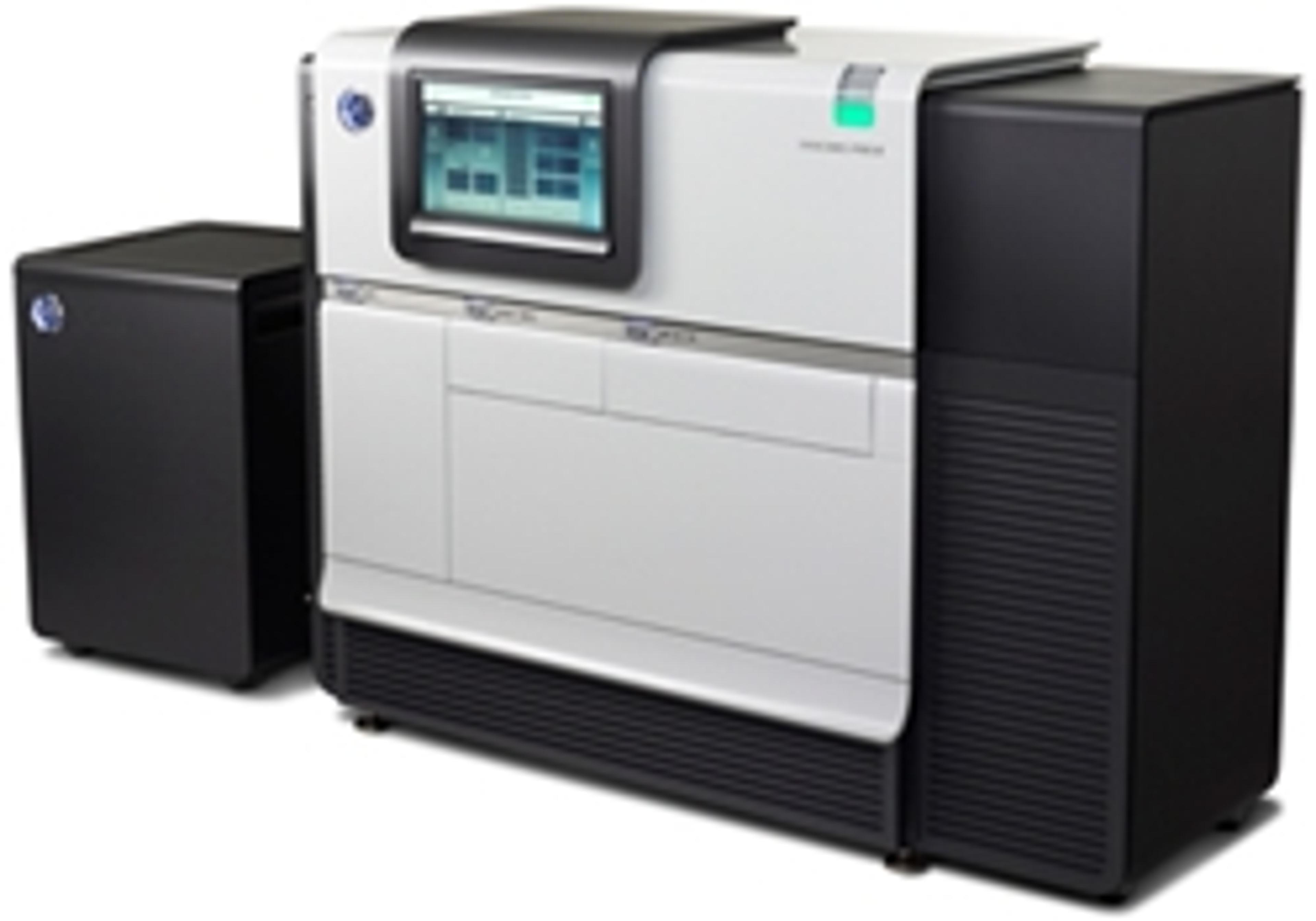
The PacBio RS II is a Single Molecule, Real-Time (SMRT®) DNA Sequencing System that provides the highest consensus accuracy and longest read lengths of any available sequencing technology. SMRT Sequencing is ideal for de novo assembly, characterization of genetic variation, methylation analysis, microbiology studies, and more.The instrument features high performance optics, automated liquid handling, and an environmental cont…

The supplier does not provide quotations for this product through SelectScience. You can search for similar products in our Product Directory.
Direct protocol, simple machine, versatile software, more than one publishable output
Sequencing of DNA samples
With the long-read sequences generated from the machine, I am able to get the complete genomic sequences of my bacterial genome and also the DNA methylation profile of my samples. The library prep is straight-forward and the sequencing part is automated. With its Iso-Seq protocol, I have a longer read thus a higher level of confidence in analyzing my sample. Besides, I can perform a hybrid assembly, using PacBio long reads as the scaffold and Illumina short reads for its accuracy and coverage. With this, I able to sequence a huge area of the repetitive region of the eukaryotic genome. Users can use the PacBio online interface for data transfer and data analysis. The online analysis portal is easy to use, and there is even a community-based forum for PacBio so that the users can share protocols. The consumables are relatively more expensive but the amount of information I can obtain from the sequences upon analysis is very helpful for my research.
Review Date: 17 Jan 2017 | Pacific Biosciences
Definitely recommend PacBio to anyone looking at small genomes,
DNA sequencing
Occasionally hits a stumbling block, but most of the time the instrument runs like a dream with little optimization required. Results are highly accurate with average read lengths over 16kB and the longest reads over 60kB consistently. Lives up to its promises. Definitely recommend PacBio to anyone looking at small genomes, repetitive regions, high GC content. I'm sure the Sequel will perform just as well.
Review Date: 23 Jun 2016 | Pacific Biosciences
NG Sequencing service
Sample preparation was a bit difficult it took quite long until results were ready.
Review Date: 30 Jan 2015 | Pacific Biosciences
DNA Sequencing High Throughput
It is convenient to use. It high read lengths with increased and high accuracy. It reads repeat regions and gives real time sequencing. It gives map-able data for use without having to sort out the quality data; hence the people with less bioinformatics capabilities can make their research ideas on sequencing into reality and by investing lesser time than PCR based sequencing systems.
Review Date: 22 Apr 2014 | Pacific Biosciences
The PacBio RS II is a Single Molecule, Real-Time (SMRT®) DNA Sequencing System that provides the highest consensus accuracy and longest read lengths of any available sequencing technology.
SMRT Sequencing is ideal for de novo assembly, characterization of genetic variation, methylation analysis, microbiology studies, and more.
The instrument features high performance optics, automated liquid handling, and an environmental control center, all directed through an intuitive touchscreen interface. The computational brain responsible for primary data analysis, called the Blade Center, is also included. This allows for seamless integration of performance enhancements through chemistry and software advances.
Features:
- Intuitive run setup tools
- Workflow optimization tools
- Error-proof instrument loading
- Robotic workflow management
- Run size flexibility – from 1 to 16 SMRT Cells per run
Applications:
- Genome finishing
- Epigenetics
- Haplotype phasing
- Repeat expansions
- Isoform sequencing
- Minor variants
Azenta Life Sciences expands fleet of PacBio HiFi sequencing systems
PacBio’s Sequel IIe devices will support genomic researchers around the world
Pacific Biosciences and Invitae to develop whole genome sequencing-based assays for pediatric epilepsy diagnostics
The collaboration will focus on the investigation of clinically relevant molecular targets for use in the development of advanced diagnostic testing for epilepsy