KAPA RNA Library Preparation Kits for Illumina
KAPA Stranded mRNA-Seq Kits includes all the enzymes and buffers required for cDNA library preparation for Illumina Next-Generation Sequencing, utilizing 100 ng – 4 µg of total RNA. KAPA mRNA Capture Beads are included for isolation of poly(A)-tailed RNA. Kits provides precise measurement of strand orientation (>99%), uniform coverage, and high-confidence mapping of alternate transcripts, and are optimized for the improved…

The supplier does not provide quotations for this product through SelectScience. You can search for similar products in our Product Directory.
KAPA Stranded mRNA-Seq Kits includes all the enzymes and buffers required for cDNA library preparation for Illumina Next-Generation Sequencing, utilizing 100 ng – 4 µg of total RNA.
KAPA mRNA Capture Beads are included for isolation of poly(A)-tailed RNA. Kits provides precise measurement of strand orientation (>99%), uniform coverage, and high-confidence mapping of alternate transcripts, and are optimized for the improved coverage of GC-rich and low-abundance transcripts. Kits contain KAPA HiFi for high-efficiency and low bias library amplification, as well as KAPA mRNA Capture Beads and a streamlined, “with-bead” protocol.
KAPA Stranded RNA-Seq Kits include all the enzymes and buffers required for cDNA library preparation for Illumina Next-Generation Sequencing, but do not contain the KAPA mRNA Capture Beads. Kits can be used to prepare libraries from 10-400 ng of either poly(A)-selected, ribosomally-depleted, or total RNA.
Features:
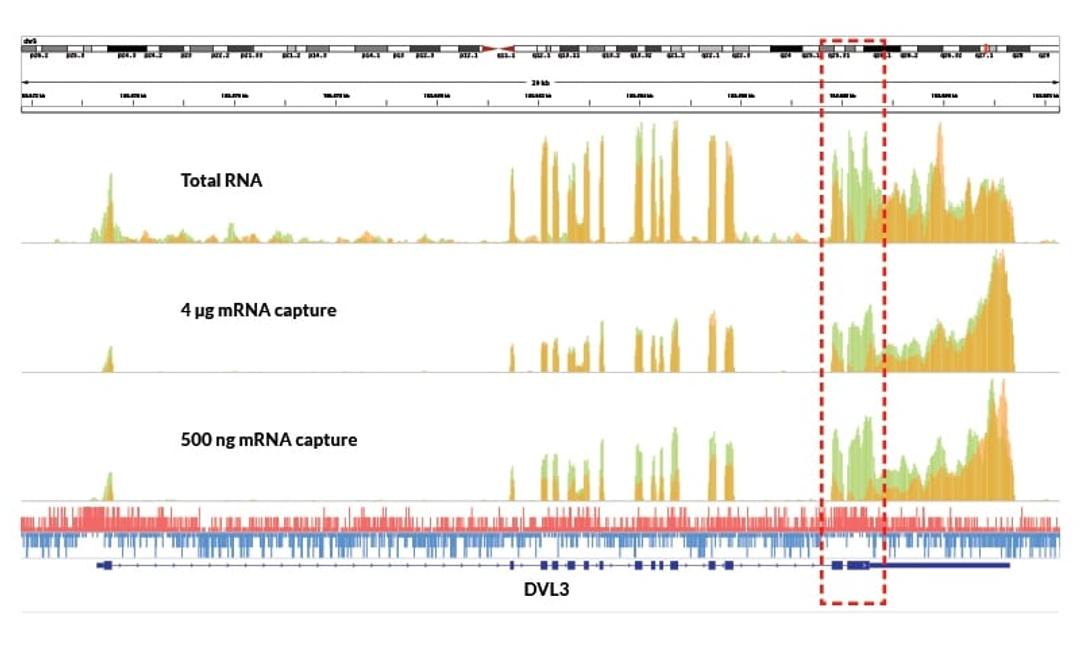
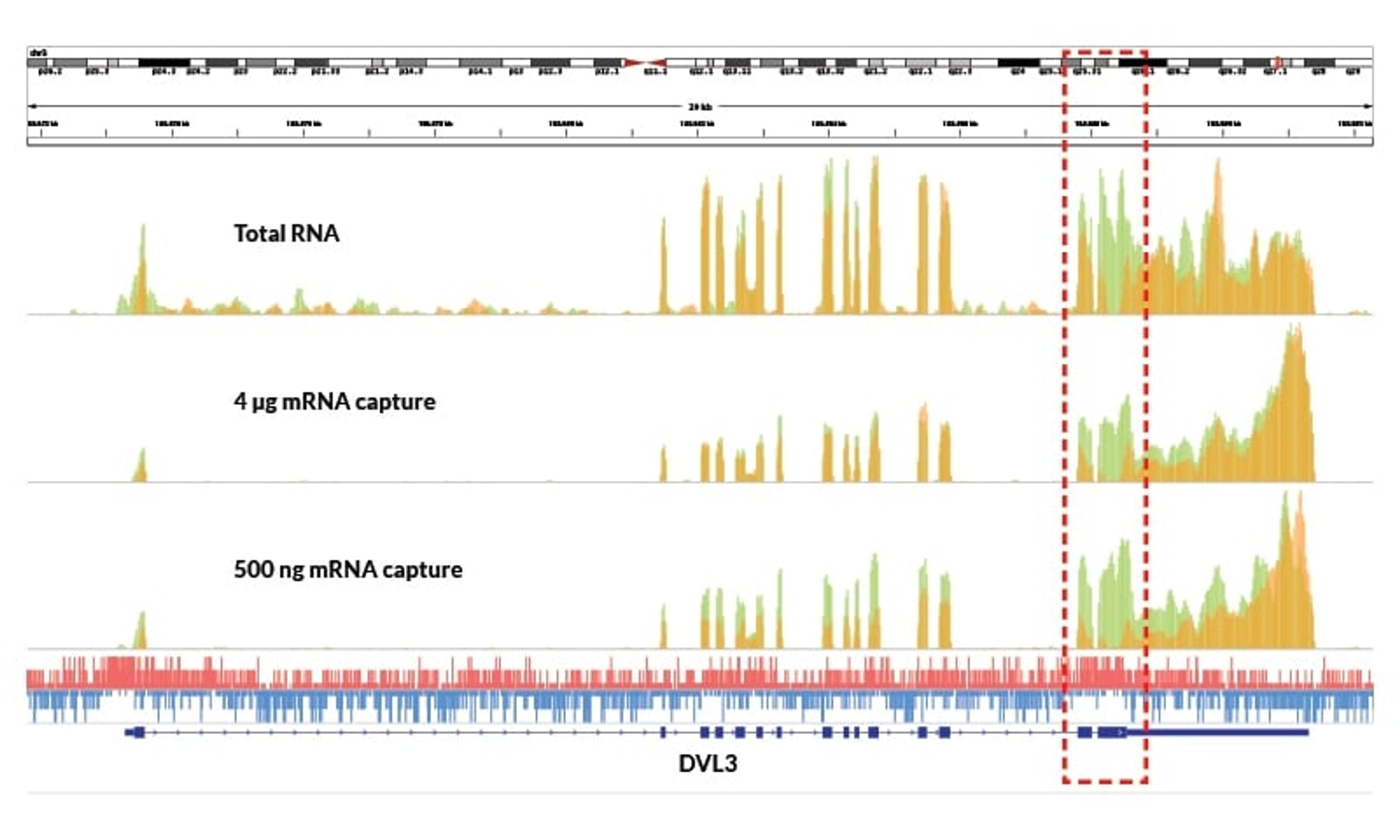
Uncover challenging transcripts
- Improved coverage of GC-rich transcripts
- Enhanced identification of exonic regions
Detect low-abundance transcripts
- Enables identification of transcripts missed by competitor kits, even with high input
- High uniformity across varying amounts of sample input
Identify more genes
- Higher percentage of uniquely mapped reads compared to Illumina TruSeq™ Stranded mRNA Sample Prep Kits
- Lower duplication rates yield better coverage
Maintain high coverage uniformity
- Minimal 5′–3′ bias across transcripts
- More uniform distribution of reads over each transcript
Applications:
- Gene expression
- Single nucleotide variation (SNV) discovery
- Post-transcriptional SNVs
- Fusion gene identification
- Targeted transcriptome
- Whole transcriptome
Automation of ChIP-Seq Library Preparation for Next Generation Sequencing on the epMotion<sup>®</sup> 5075 TMX
This application note describes the successful automation of Illumina ChIP-Seq library preparation on the Eppendorf epMotion® 5075 TMX, using the KAPA High-Throughput Library Preparation Kit from Kapa Biosystems. ChIP-Seq library preparation can be a challenging procedure to automate because of its low ChIP DNA input. The combined advantages of the Eppendorf epMotion® 5075 TMX and the KAPA High-Throughput Library Preparation Kit offer an industry-leading solution for standard and low-input library preparation, particularly for ChIP-Seq.
Co-optimization of Probes and Polymerases to Drive the Evolution of Targeted Sequencing
This poster demonstrates the capabilities of targeted sequence enrichment using the KAPA HTP Library Preparation Kit and KAPA HiFi HotStart DNA Polymerase. Inefficiencies in library construction and automation-unfriendly workflows have previously meant low throughput and poor performance. By combining enzymes specifically tailored for high performance NGS applications with innovative oligonucleotide probe design, improved library construction and targeted enrichment can be achieved.
An Effective and Reliable Enzymatic Method for the Depletion of Ribosomal RNA for Preparation of High-Quality Libraries for Transcriptome Sequencing
Ribosomal RNA (rRNA) accounts for the vast majority of total cellular RNA, and its efficient removal is critical during the RNA-Seq library preparation process in order to maximize the yield of biologically-informative transcriptome-derived reads. Sequencing of total RNA samples that have been depleted of rRNA provides a more comprehensive representation of the transcriptome compared to mRNA sequencing. This application note demonstrates a flexible and highly effective method for targeted enzymatic depletion of rRNA using complementary oligonucleotide probes and RNase H degradation.
Application of NGS in the Diagnostic Lab on Archival FFPE Tissues
Hosted by Kapa Biosystems at AMP2015, Dr. Brian Walker of the Royal Marsden Hospital presents at a work shop entitled "Application of NGS in the diagnostic lab on archival FFPE tissues".