Seven Novel PCR Applications
23 Apr 2015Editorial article

Improve your PCR methods with this round-up of recent applications
Discover new ideas for your PCR applications with these seven recently published application notes and methods. Or, if you haven’t found the application you require, search our extensive application note library to find the method you need. There are more than 10,000 methods and applications to download for free!
1. Intercalating Fluorescence Dyes: A Comparison of Different Kits Using the qTOWER from Analytik Jena
Using intercalating dyes for quantification assays is one of the most common and popular real-time PCR applications. SYBR® Green or EvaGreen® for example are easy to use. Thus a wide variety of ready-to-use master mixes are available to simplify workflows as much as possible. Analytik Jena’s standard real-time thermal cycler qTOWER 2.0/2.2 was used to compare the performance of those mixes.
2. Droplet Digital™ PCR: Multiplex Detection of KRAS Mutations in Formalin-Fixed, Paraffin-Embedded Colorectal Cancer Samples
KRAS is mutated in approximately 40% of colorectal cancers. To optimize therapy strategies for personalized care, it is critical to rapidly screen patient samples for the presence of multiple KRAS mutations. This application note demonstrates a multiplexing strategy which has been developed to screen seven actionable KRAS mutations in colorectal cancer samples using digital PCR. This sensitive and inexpensive method reduces the risk of contamination and can be easily implemented for rapid, routine screening of cancer samples.
3. Discriminating Copy Number Variation from Heterogeneous Samples Using the S3™ Cell Sorter and the QX200™ Droplet Digital™ PCR System
This application note shows that combining technologies such as flow cytometry, cell sorting, and digital PCR can enable highly accurate and more specific results attributed to cell populations of interest vs. the bulk heterogeneous populations. By pairing tools like Bio-Rad’s S3 Cell Sorter and the QX200 Droplet Digital PCR System, targeted populations can be further characterized to uncover correlations in cancers and diseases.
4. Xpose Reader App: Quantification of Purified PCR Samples with Spectral Content Profiling
An app for specific quantification of DNA amplicon in purified PCR samples is available on the Xpose ‘Touch & Go’ reader. Trinean’s proprietary spectral content profiling is used to isolate the profile of the desired molecule (here DNA) from the measured UV/VIS spectrum, thereby distinguishing it from absorbing sample contaminants. The specific DNA profile is then used to determine the DNA concentration. This application note describes how to use this application and how to interpret its results.
5. Precise Evaluation of Plant RNA Extraction Methods with Automated RT-qPCR Assay
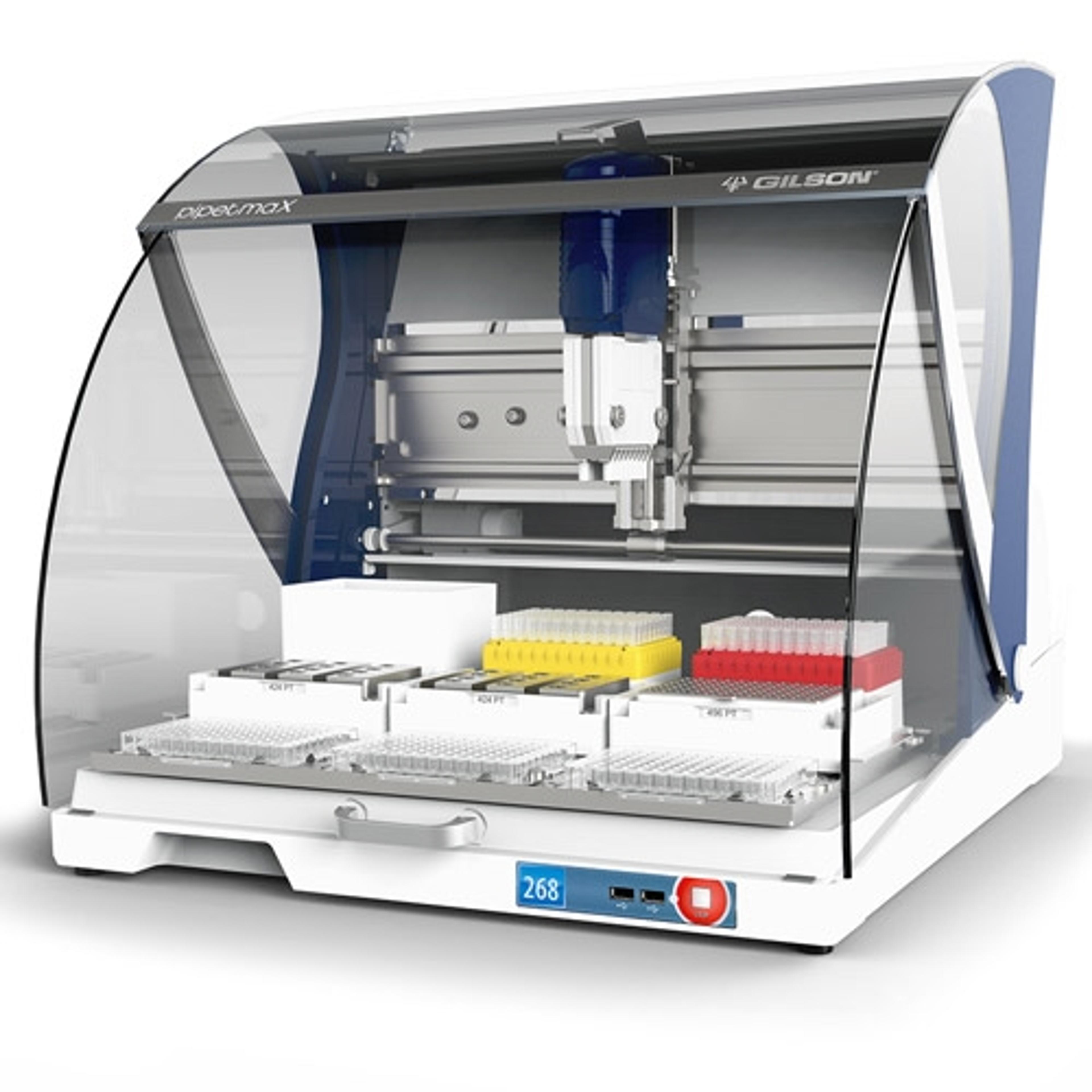
In this application note, functional corn RNA was quantitated by RT-qPCR using the GoTaq® Probe 1-Step RT-qPCR System after two different extraction methods: Promega’s Maxwell® 16 LEV (low elution volume) Plant RNA Kit and a competitor spin column-based plant RNA extraction kit. To enhance the pipetting accuracy and consistency, an automated pipetting assistant, Gilson PIPETMAX® 268, was used to reduce pipet-ting errors inherent in manual methods.
6. Thermal Shift Assay Using SYPRO® Orange and qTOWER 2.2 to Detect Protein Melting Temperatures
Protein thermal shift assays are sensitive and rapid tools to examine protein thermal stability, helping to evaluate protein-ligand binding and thus to finding optimal buffer conditions or analyzing protein variations. This application note demonstrates the generation of a melting curve using the qTOWER 2.2 real-time PCR thermal cycler. This allows an easy and fast determination of protein melting temperatures.
7. Performance of Axygen® Maxymum Recovery® Pipet Tips
With the emergence of high-throughput PCR and DNA sequencing, reliable sample preparation and delivery methods have become a critical issue. Variations introduced as a result of pipet tip performance are often far greater than 1% of the target value and overall performance can be seriously compromised. This application note demonstrates the performance of Axygen Maxymum Recovery pipet tips, which have been specifically designed for applications requiring high accuracy and reproducibility.