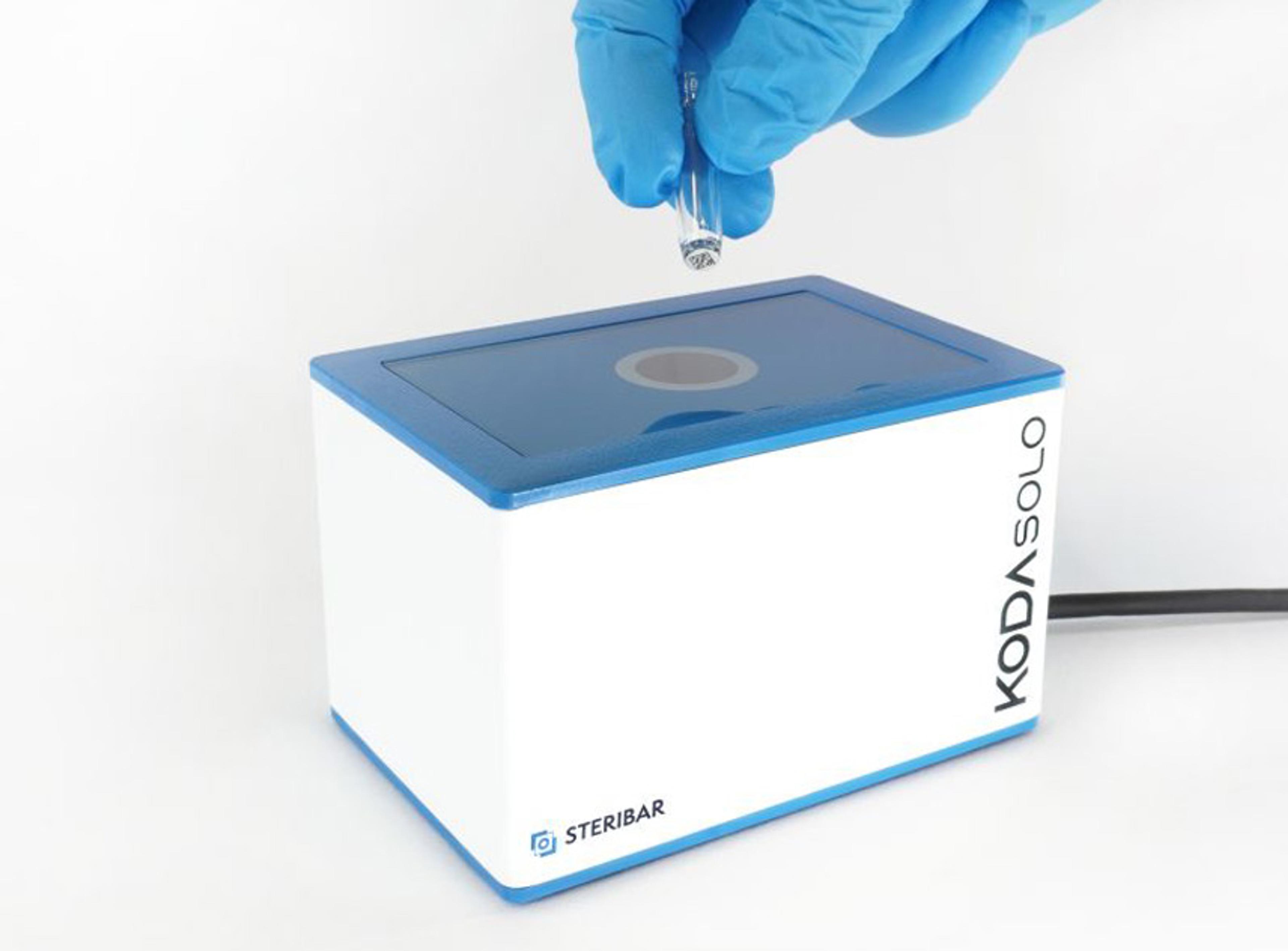
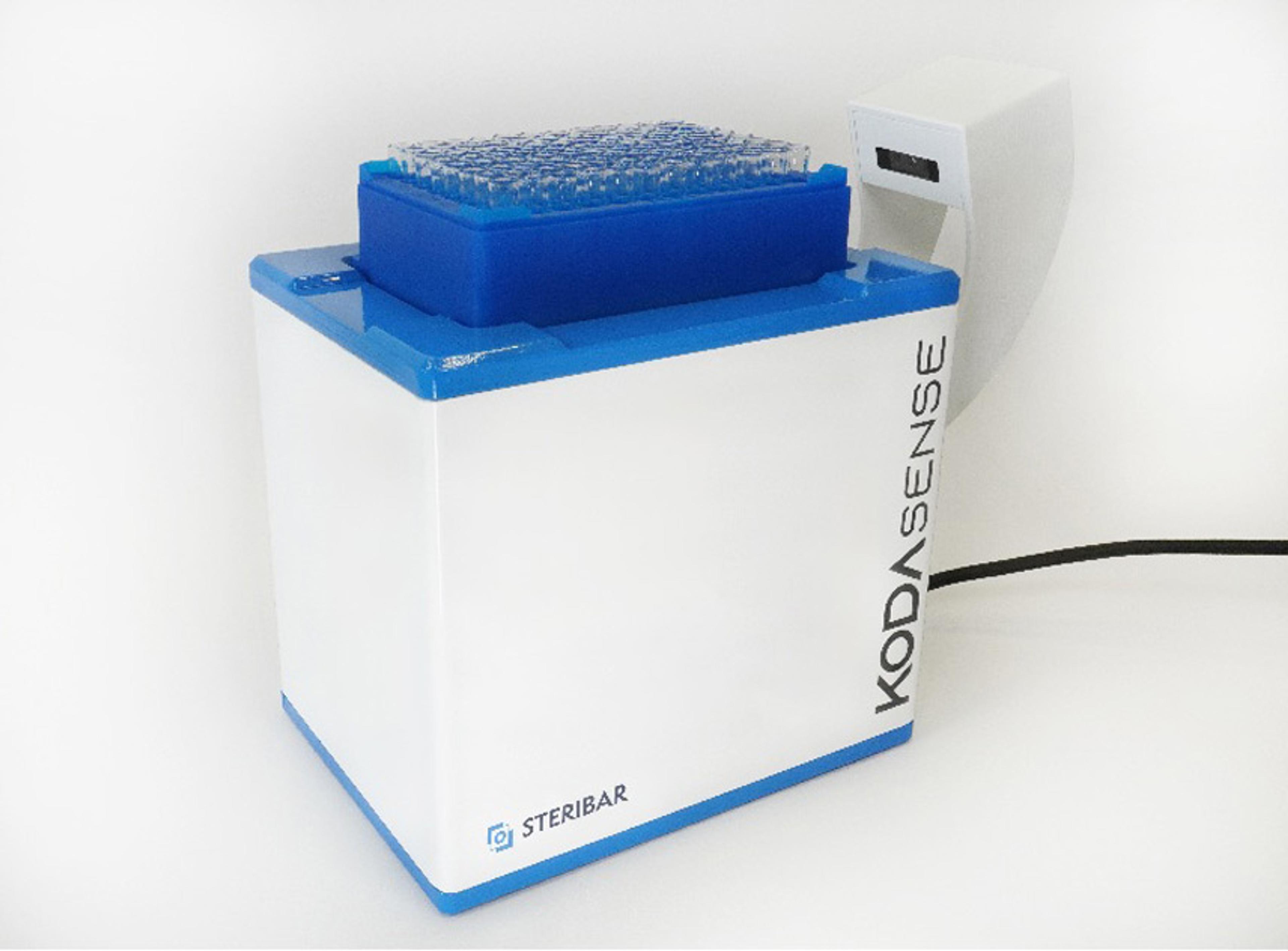
PhenoImager® Fusion
The PhenoImager ® ; Fusion system is a whole-slide, single-cell resolution, ultrafast multispectral imager ideal for standard throughput brightfield and fluorescence imaging applications. This imaging platform can be integrated with the PhenoCycler ® ; system (formerly CODEX ® ) for ultra high-plex spatial discovery at scale.

The supplier does not provide quotations for this product through SelectScience. You can search for similar products in our Product Directory.
Great experience thus far
Spatial omics of brain tissue
We are developing CNN model to segment neurons and astrocytes, etc
Review Date: 26 Sept 2022 | Akoya Biosciences
Easy conventional platform to identify cell types and the levels of target molecule.
Analyze immune cell activities in human and mouse derived tissue samples
Fluorescence based multispectral imaging can be performed with Vectra. We have used Vectra for more than 3 years, and analyzed any tissue samples stained for multiplex IHC. This imaging platform can be a good solution for detecting up to 7 markers. It can also scan slides stained with H&E or chromogenic IHC. We have tested detection limit for fluorescence signal. If the signal is weak (e.g. immunofluorescence without signal amplification like immunohistochemistry), it may or may not detect the signal because of the limited exposure time for scanning. Immunohistochemically amplified signals usually do not have problem being detected. However, when the signal is super high on one marker, it may disrupt unmixing procedure on inForm software, so the staining conditions need to be optimized to acquire signals from multiple markers. Scanning slides is easy to perform, and it does not need to be in the dark room. Scanning speed is quite fast to acquire entire tissue image (x4 and x10) and multi-spectral images (x20, x40). Analysis for scanned images is easily performed on inForm software. The inForm software can do: (1) unmix the each fluorescence marker signal from multispectral image, (2) perform tissue segmentation (if tissue marker like cytokeratin is stained), (3) cell segmentation to locate each cell location on the image and fractionate subcellular compartment (nucleus, cytoplasm and membrane), (4) score marker levels and (5) summarize and convert dataset into tissue and cell related parameters and different image format. The parameters indicating the levels of each marker can be used for FACS like analysis. Converted images into multilayered TIFF file can be opened with other imaging software. This AI supported imaging analysis procedure with inForm is getting easier in newer version. However, if tissue architecture is unique (e.g. cells are elongated, distance of nucleus to membrane is big, the tissue has various size of nuclei), it may take much time to train the AI or it may not clearly show each cell membrane. The maintenance is minimal like a conventional microscope. Since a light bulb for detecting fluorescence needs to be replaced after a certain number of hours, we have upgraded the light source to LED. The worst thing on Vectra is actually when the light bulb is reaching a certain time. It will keep beeping, so no one wants to be in the room! Customer care for technical service is very quick and user friendly. As a scientist who has been working with various microscopes, it is not good enough to acquire molecular localization on a particular organelles but this Vectra system is good for identifying cell types in tissue sample and it is also good for quantifying the levels of markers in each cell in the tissue sample.
Review Date: 3 Dec 2020 | Akoya Biosciences
Capable of both brightfield and fluorescence whole-slide imaging, the PhenoImager Fusion instruments serves the needs of laboratories with standard throughput needs while still delivering high quality imaging with fast scanning times. Combined with our proprietary Multispectral Imaging (MSI) Technology (patent pending) which allows for easy detection and the measurement of multiple overlapping biomarkers within a single tissue without the interference of autofluorescence and fluorophore crosstalk, the PhenoImager Fusion provides you with confidence to accurately phenotype and quantify tumor-immune cell interactions in the tissue microenvironment (TME) and develop biomarker signatures with higher predictive accuracy.
Spatial phenotyping without compromise in plex, resolution, or throughput
Biological systems are complex, and full spatial understanding of tissue heterogeneity and cellular microenvironments has been hampered by trade-offs that are forced by the available analytical technologies.
All of which has given rise to the concept of the ‘iron triangle’ – the hard problem of improving resolution, plex and throughput simultaneously in one analytical solution.
But the world is set to change with the advent of new spatial phenotyping approaches that enable viewing, characterization and quantification of cells by lineage and variant with single-cell resolution, all in the context of an intact tissue sample.
From capturing tumor heterogeneity and validating cancer models, to teasing out how the cellular microenvironment impacts disease progression and treatment response, this guide will explain how spatial phenotyping can deliver advances in automation, efficiency, tunability, and speed that are set to transform how we study tissue biology.
Next-generation cancer treatment: Patient stratification with immuno-oncology
This video explains how using biomarkers on tissue samples not only provides new insights into the biology of cancer but ultimately enables a new level of patient stratification. This allows the most effective treatment to be used for any given cancer leading to significantly improved outcomes for patients.
Akoya Biosciences launches PhenoCode Signature Panels to accelerate development of predictive biomarkers for cancer immunotherapy
Each panel focuses on distinct areas of tumor biology and response to therapy which are of greatest interest to translational and clinical researchers
9 top new resources for your biopharmaceuticals research
Exclusive interviews, new methods, free downloads and much more to help advance your biopharmaceutical research