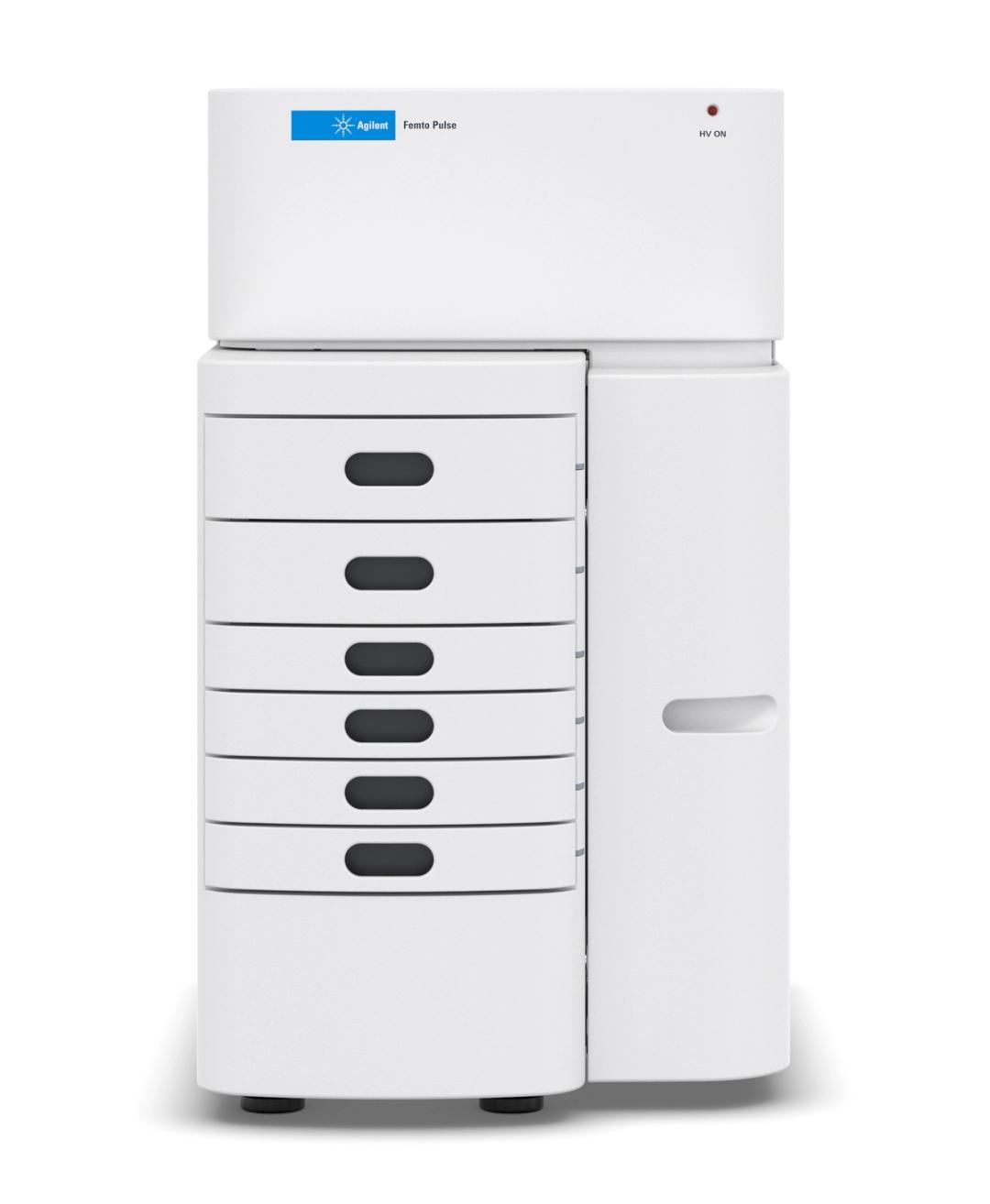
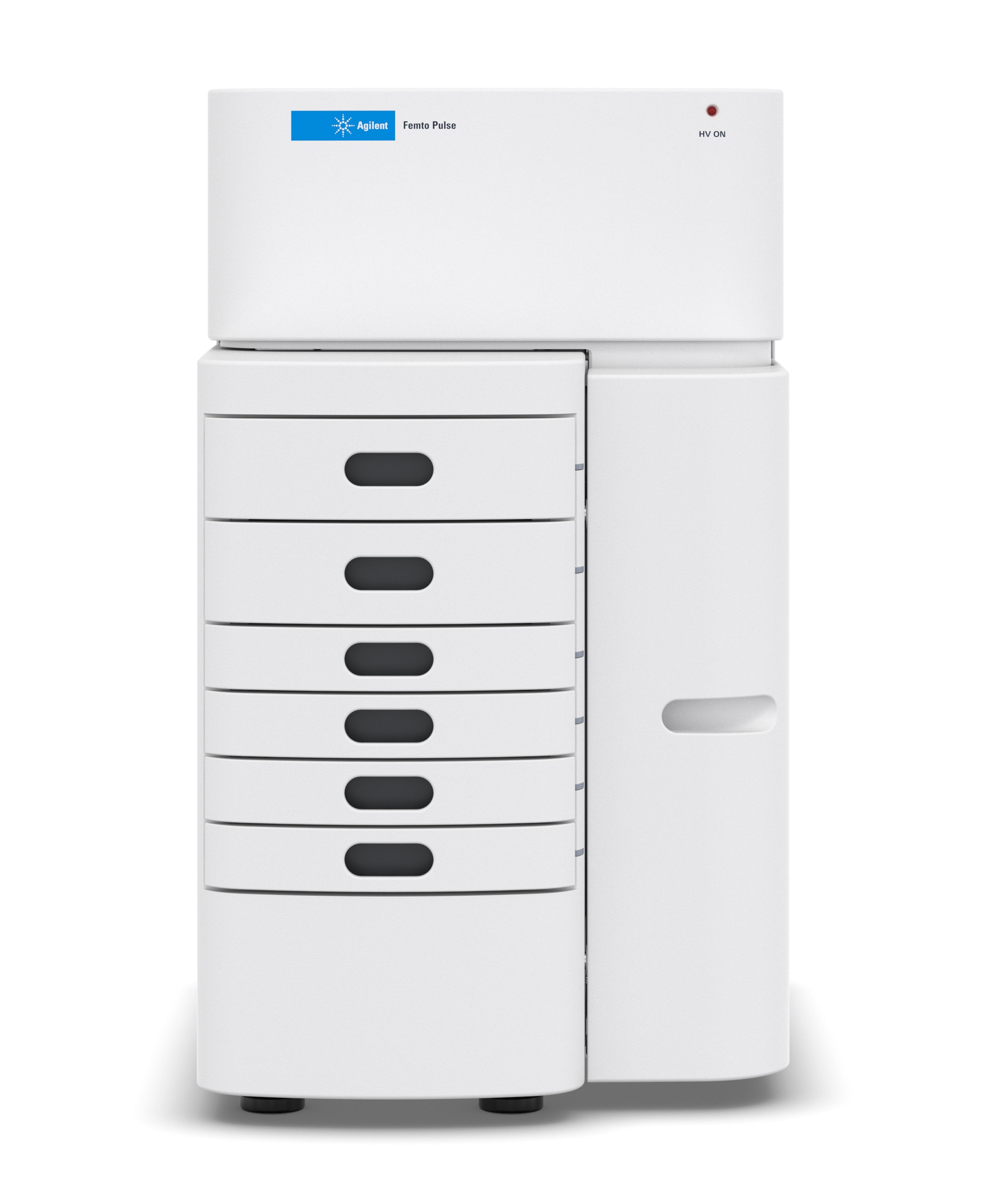
Femto Pulse System
The Agilent Femto Pulse system delivers remarkable sizing and sensitivity to researchers.
Shatter analytic barriers with pulsed-field capillary electrophoresis. The Agilent Femto Pulse system provides an intuitive platform for researchers to analyze low-concentration, high molecular weight nucleic acid samples quickly and accurately.
Confidently size DNA smears through 165 kb and DNA fragments through 200 kb in approximately one and two hours, respectively. Compared to legacy methods for the analysis of high molecular weight DNA samples, the Femto Pulse completes separations up to 20x faster, accelerating workflows and putting crucial results in the hands of researchers sooner. Additionally, the Femto Pulse provides unmatched sensitivity, detecting DNA fragments down to 5 fg/µL (in-well concentration).
Features & benefits:
- Femtogram-level sensitivity – Conserve precious sample with up to 10x higher sensitivity for nucleic acid smears, and up to 100x higher sensitivity for nucleic acid fragments compared to other analytical instruments, and 1000x greater sensitivity than PFGE.
- Streamlined sample preparation – Minimal sample handling reduces the risk for sample degradation or contamination.
- Large smear resolution – Size DNA smears through 165 Kb in about 1 hour with automated operation and analysis.
Brochures
Agilent automated electrophoresis portfolio brochure
Whether performing low- or ultra-high-throughput testing, Agilent Technologies' automated electrophoresis instrument portfolio delivers reliable, objective assessments of sample integrity, concentration, and fragment size. Explore the complete portfolio of automated electrophoresis instruments, application-specific assays, intuitive software, and expert technical services that provide flexible solutions for robust and objective sample quality assessment.
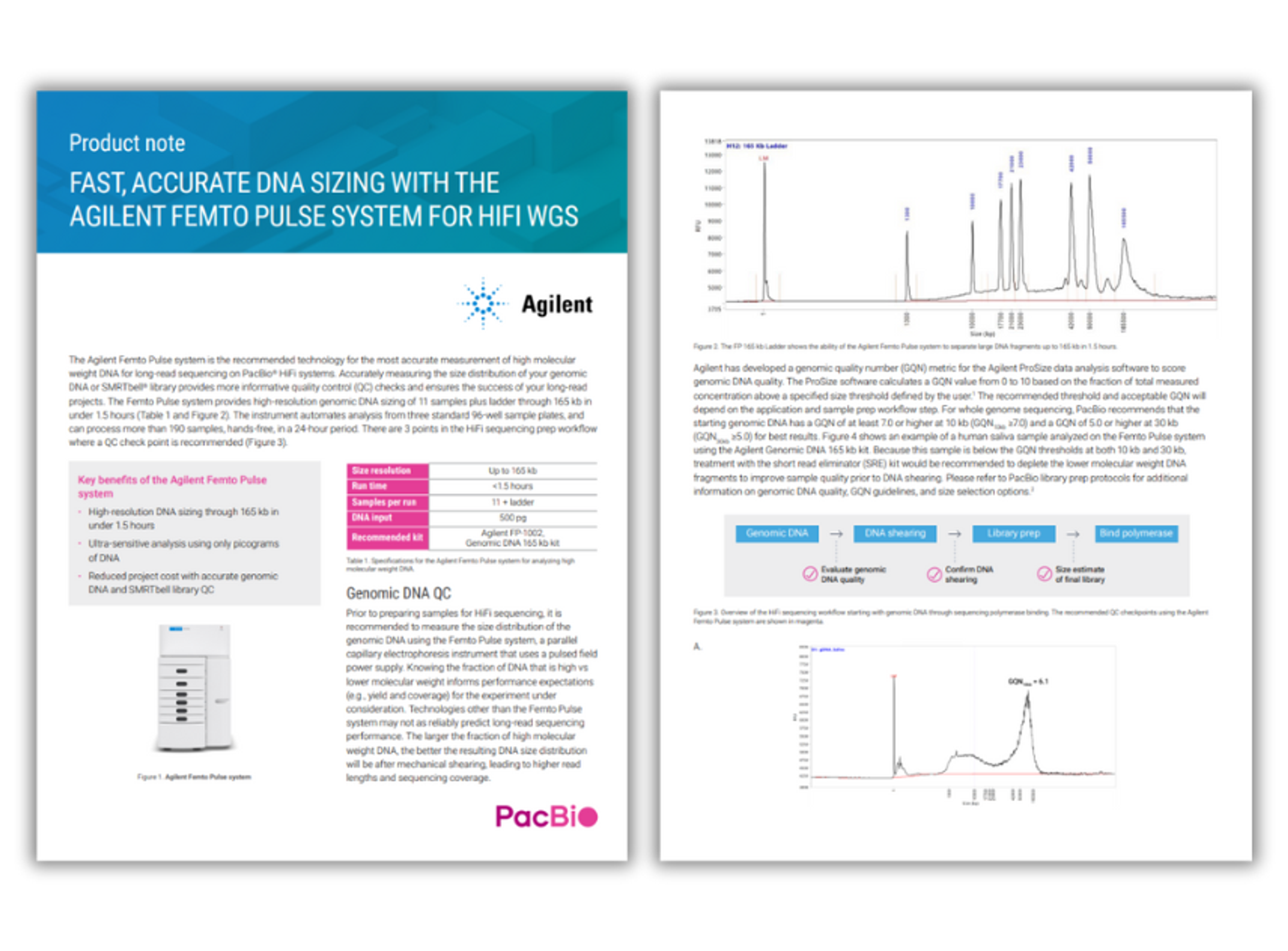
Fast, accurate DNA sizing with the Agilent Femto Pulse system for HIFI WGS
Achieve fast, accurate sizing of high-molecular-weight DNA for PacBio® HiFi whole-genome sequencing using the Agilent Femto Pulse system. This ultra-sensitive pulsed-field capillary electrophoresis platform enables precise genomic DNA and SMRTbell® library QC, ensuring optimal read length, yield, and sequencing performance from as little as picogram DNA input.
Quality control in the PacBio HiFi whole genome sequencing library preparation workflow
Accurate and reliable next-generation sequencing (NGS) results rely on starting with high-quality samples. In long-read sequencing workflows, such as the PacBio HiFi workflow, essential quality control (QC) steps include assessing input gDNA, intermediate steps throughout library preparation, and the final library. Electrophoretic analysis enables evaluation of sample size, purity, and integrity at each of these steps. Agilent offers several automated electrophoresis systems, including the Agilent Femto Pulse, Fragment Analyzer, and TapeStation systems, each with unique benefits for DNA QC.
This application note demonstrates the capabilities and limitations of each of the automated electrophoresis systems within the PacBio HiFi whole genome sequencing (WGS) library preparation workflow, aiding in the selection of the appropriate QC system.
Visualizing DNA for long-read sequencing by moles, not mass
Traditional measurements during NGS library preparation include the determination of the sample mass within specific size ranges. This analysis is easily performed with Agilent automated electrophoresis instruments, which provide visual results in the form of a digital gel image and electropherogram. While this representation has been widely used for quality control of sheared gDNA and the final NGS library, examining the molarity of a sample may provide a better visual representation of the number of sequencing reads that can be produced by a sample, especially for long-read sequencing. Explore Agilent's analysis of high molecular weight samples using the Agilent Femto Pulse system and the accompanying ProSize data analysis software.
DNA shearing devices for short- and long-read sequencing
One of the main focuses of Hologic Diagenode’s epigenetics department is developing DNA and chromatin shearing devices for short- and long-read sequencing. During DNA sizing and quality analysis, the team opted to use the Agilent Femto Pulse and Fragment Analyzer systems, which can also be utilized for applications such as RNA, chromatin-based methylation studies, and NGS library profiling. Hologic Diagenode explains how its team uses these Agilent systems to achieve higher throughout and greater flexibility in its work, supporting the goal of getting new devices to customers.
Genomic DNA handling and the Agilent Femto Pulse system
In this application note, Agilent Technologies discusses the significance of genomic DNA (gDNA) integrity in molecular biology labs. Maintaining high-quality samples is crucial to generating reliable results and saving time and resources. Various factors such as extraction techniques, storage conditions, and mixing can affect gDNA sample integrity. Pulsed field gel electrophoresis (PFGE) is commonly used for quality assessment but requires significant sample quantities and overnight runs. Agilent Femto Pulse system with Genomic DNA 165 kb kit is a faster and cost-effective alternative that offers high sensitivity and resolution. This note examines the impact of different handling protocols on gDNA sample integrity and provides recommendations for best practices when using the Femto Pulse.
Fast separation, ultra sensitivity, one instrument
The Agilent Femto Pulse system separates high molecular weight (HMW) DNA fragments and detects nucleic acids into the femtogram range from low concentration samples. Download this application note to find out more.
Comparison of Agilent Femto Pulse System sizing with long-read sequencing read length
This application note by Agilent demonstrates how the Femto Pulse was used in a long-read sequencing workflow.
Benefits of the Femto Pulse System versus PFGE
This application note by Agilent displays the benefits of the Femto Pulse System versus PFGE.
Genomic DNA handling and the Agilent Femto Pulse System
This application note explores the impact that different handling protocols can have on gDNA sample integrity and provides recommendations for best practices when performing analysis with the Femto Pulse.
Is Sample Integrity Assessment for Long-Read Sequencing Technologies Essential?
Exploring Key Topics in Sample Quality Control for NGS Workflows
Next-generation sequencing (NGS) technologies enable deeper insights into different areas of research. In this latest webinar series, researchers from the European Molecular Biology Laboratory (EMBL), Quantabio LLC,, and Agilent Technologies discuss the importance of input sample quality control in whole genome sequencing (WGS), panel-based targeted sequencing and long-read sequencing workflows in order to maximize sequencing output.
Sample Integrity Assessment for Long-read Sequencing Technologies, Is It Essential?
Next generation sequencing (NGS) has evolved rapidly in the past decade creating opportunities for new technologies to emerge, ultimately making this technology more robust. Among these, particularly long-read sequencing has experienced an increase of popularity in the research community due to improvements in the overall stability and the quality of generated data.
However, the value of long-read sequencing data is critically dependent on the quality of the input material used for a library preparation. Therefore, reliable and accurate quality control (QC) is essential for the success of these sequencing technologies. At EMBL, the Agilent Femto Pulse system is a fundamental part of the daily routines for processing LRS samples. It allows researchers to verify the integrity and fragment size distribution of nucleic acid samples.
This webinar will highlight the importance of Femto Pulse in GeneCore workflows and present data that emphasize the significance of sample quality control for success of EMBL’s undertakings.
Key learning objectives:
- How stringent QC of incoming samples helps identify and prevent potential problems that could negatively impact their performance in library preparation and sequencing
- How ensuring the accurate size and quantity of the finished library is essential for maximizing sequencing output.
- How the Agilent Femto Pulse system is used in GeneCore workflows.
Who should attend:
- NGS and genomics researchers
- Molecular biologists
- Cancer researchers
- Sequencing lab managers and technicians
Certificate of attendance
- All webinar participants can request a certificate of attendance, including a learning outcomes summary, for continuing education purposes.
Streamlined DNA Library Preparation Solutions and Guidelines for Robust Quality Control of PCR-free WGS Libraries
Exploring Key Topics in Sample Quality Control for NGS Workflows
Next-generation sequencing (NGS) technologies enable deeper insights into different areas of research. In this latest webinar series, researchers from the European Molecular Biology Laboratory (EMBL), Quantabio LLC,, and Agilent Technologies discuss the importance of input sample quality control in whole genome sequencing (WGS), panel-based targeted sequencing and long-read sequencing workflows in order to maximize sequencing output.
Streamlined DNA Library Preparation Solutions and Guidelines for Robust Quality Control of PCR-free WGS Libraries
Whole genome sequencing (WGS) and panel-based targeted sequencing provide vital genetic information for various applications. The quality of sequencing data hinges on the input library, traditionally prepared using PCR amplification. PCR-free WGS library preparation has become a critical alternative, eliminating amplification-induced issues and preserving the genome's true representation. This is especially important for accurate mutational analysis and disease diagnostics, enhancing the detection of structural variants and GC-rich regions. Quality control (QC) of PCR-free libraries presents a challenge when using electrophoresis. This typically results in inaccurate library size estimations due to the structural features of Y-shaped adapters commonly used during library preparation. In this webinar, we describe a method to accurately determine the size of PCR-free libraries by adding a short PCR amplification step prior to automated DNA fragment analysis.
Key learning objectives:
- Understand why PCR-free WGS library preparation is essential for improving the reliability and efficiency of genetic analyses across various fields
- Discover how sparQ DNA Frag and DNA Library Prep Kits can be utilized to generate PCR-free libraries for DNA library preparation
- Learn how to modify PCR-free libraries for proper QC utilizing the Agilent TapeStation system
Who should attend:
- NGS and genomics researchers
- Molecular biologists
- Cancer researchers
- Sequencing lab managers and technicians
Certificate of attendance
- All webinar participants can request a certificate of attendance, including a learning outcomes summary, for continuing education purposes.
How automation and integrated sample QC enhance a targeted long-read-sequencing workflow
Tuesday, November 4, at 16:00 GMT | 17:00 CET | 11:00 EST | 8:00 PST
As the demand for greater accuracy and efficiency in genetic analysis grows, new approaches are needed to overcome the blind spots of conventional sequencing methods. This webinar will showcase a fully automatable workflow that integrates Agilent SureSelect hybrid capture, Agilent automated electrophoresis systems for sample quality control with PacBio HiFi long-read sequencing.
In addition to discussing workflow, we’ll share case studies where long reads play a fundamental role in identifying genetic variants that may be missed by conventional sequencing in research studies.
Join us to learn how targeted long-read sequencing:
- Focuses on genes and regions of interest in genetic research, while reducing costs and boosting sequencing depth at key loci.
- Overcomes the limitations of short-read technologies, including challenges with pseudogenes, homologous loci, structural variants, complex transcript isoforms, splicing events, and difficult regions such as repetitive or GC-rich sequences.
- Enhances both analytical performance and laboratory efficiency through automation.
Key learning objectives:
- Gain practical insights into implementing an automated targeted long-read sequencing workflow in the laboratory.
- Identify the genetic mutation types best suited for targeted long-read sequencing approaches.
- Understand the analytical advantages of long-read sequencing compared with conventional short-read methods.
Who should attend?
Lab directors, molecular pathologists, research scientists, clinical lab technicians, and anyone running or interested in running NGS workflows.
Certificate of attendance
If you attend the live webinar, you will automatically receive a certificate of attendance, including a learning outcomes summary, for continuing education purposes.
If you view the on-demand webinar, you can request a certificate of attendance by emailing editor@selectscience.net.
For Research Use Only. Not for use in diagnostic procedures. PR7003-294
Simplified sample preparation for targeted Next-Generation Sequencing
Next-generation sequencing (NGS) has a broad spectrum of laboratory applications, from life science to drug discovery. It is important to optimize every stage of your NGS workflow to ensure accurate and reliable results. Following May’s SelectScience webinars, this series explores how you can promote confidence in your NGS experiments by simplifying and accelerating NGS sample preparation and nucleic acid sample quality control.
Simplified sample preparation for targeted Next-Generation Sequencing
Targeted NGS is essential for genomic analysis, including cancer, inherited and infectious disease research. This webinar will discuss a simplified sequencing workflow that enables an optimized, hands-free process for automated sample preparation. Our integrated solutions for sample QC, library preparation and target enrichment deliver reliable sample quality for reproducible results.
Key learning objectives
- Learn how NGS sample preparation workflows can be simplified with Agilent solutions
- Learn about the importance of NGS sample QC and the Agilent instrumentation
- Learn about the SureSelect target enrichment system to deliver robust, traceable, and reproducible sequencing data
- Learn how the Magnis NGS Prep system provides a completely automated workflow for NGS library preparation with SureSelect reagents
Certificate of attendance
All webinar participants can request a certificate of attendance, including a learning outcomes summary, for continuing education purposes.
NGS sample quality control: Your questions answered
Discover the variations in different methods of isolating high-molecular weight DNA for nanopore sequencing