Swift Accel-NGS® 2S Plus DNA Library Kit (96 reactions)
Optimal sequencing of your cfDNA, FFPE and ChIP DNA

The supplier does not provide quotations for this product through SelectScience. You can search for similar products in our Product Directory.
Minimizes bias and supports inputs as low as 10 pg.
Benefits include:
- Exceptional yields from DNA inputs of 10 pg to 250 ng
- Excellent efficiency, data quality, and evenness of coverage
- High complexity libraries for deep sequencing
- No base composition bias from AT-/GC-rich genomes
This is the best choice for users working with cfDNA and degraded FFPE samples as well as conducting ChIP-Seq.
Accel-NGS® 2S DNA Library Kits
The application notes provides an overview of Accel-NGS® 2S DNA Library Kits. Accel-NGS 2S DNA Library Kits produce high quality libraries with an all-inclusive, easy-to-use format. Applications of the Library Kits, including use in sequencing liquid biopsy samples and Pico-Scale ChIP-Seq are also covered.
Understanding NGS Intricacy and Single-Cell Sequencing
The application note describes the sequence coverage performance and preservation of molecular complexity of next generation sequencing (NGS) libraries, generated from human and microbial genomic DNA using Accel-NGS® 2S DNA Library Kits for whole-genome sequencing (WGS) on the Illumina® platform. Comparisons to the leading commercially available methods are also presented.
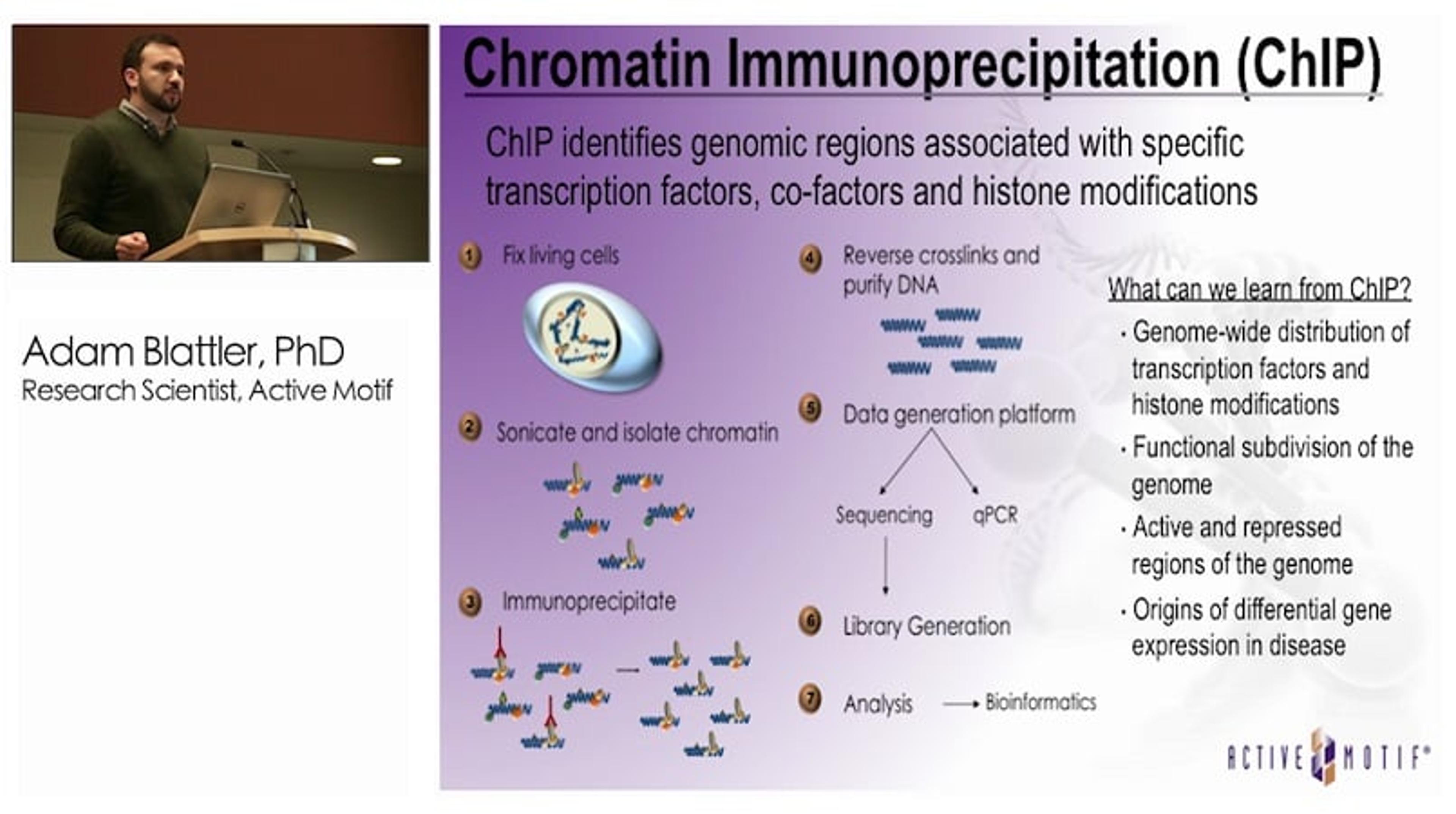
New Tools for Studying Epigenomes of Clinical Samples
Adam Blattler, a Research Scientist at Active Motif, gave an informative presentation on ChIP-Seq (Chromatin Immunoprecipitation Sequencing) and how it can be performed on FFPE (formalin-fixed, paraffin-embedded) clinical samples. Adam gives a background on epigenetic regulation and histone modification and explains the theory behind ChIP-Seq. He describes how Swift Biosciences have developed a highly sensitivity kit, the Accel-NGS® 2S Plus, which overcomes the problem of low abundance of chromatin, an issue encountered when using FFPE samples. Finally, Adam describes new advancements in ChIP-Seq technology such as enChIP (Engineered DNA binding molecule mediated ChIP), which can identify chromosomal looping.
Accurate De-Duplication of Sequencing Data with New Molecular IDs from Swift Biosciences
Kate Cunningham, PhD, Applications Scientist at Swift Biosciences, introduces the new range of molecular IDs for Accel-NGS® 2S library kits. Hear how the molecular IDs label each individual library molecules, to enable low frequency variant detection, as well as accurate de-duplication of single read sequencing and sequencing from samples with non-random fragmentation. The technology is ideal for applications including whole exome sequencing (WES), deep targeted sequencing, cell-free DNA sequencing and ChIP-Seq.
Advances in Sequencing Technology Facilitate Gene Expression Analyses at the Epigenomic Level
Three leading scientists discuss their research and the benefits of ChIP-Sequencing