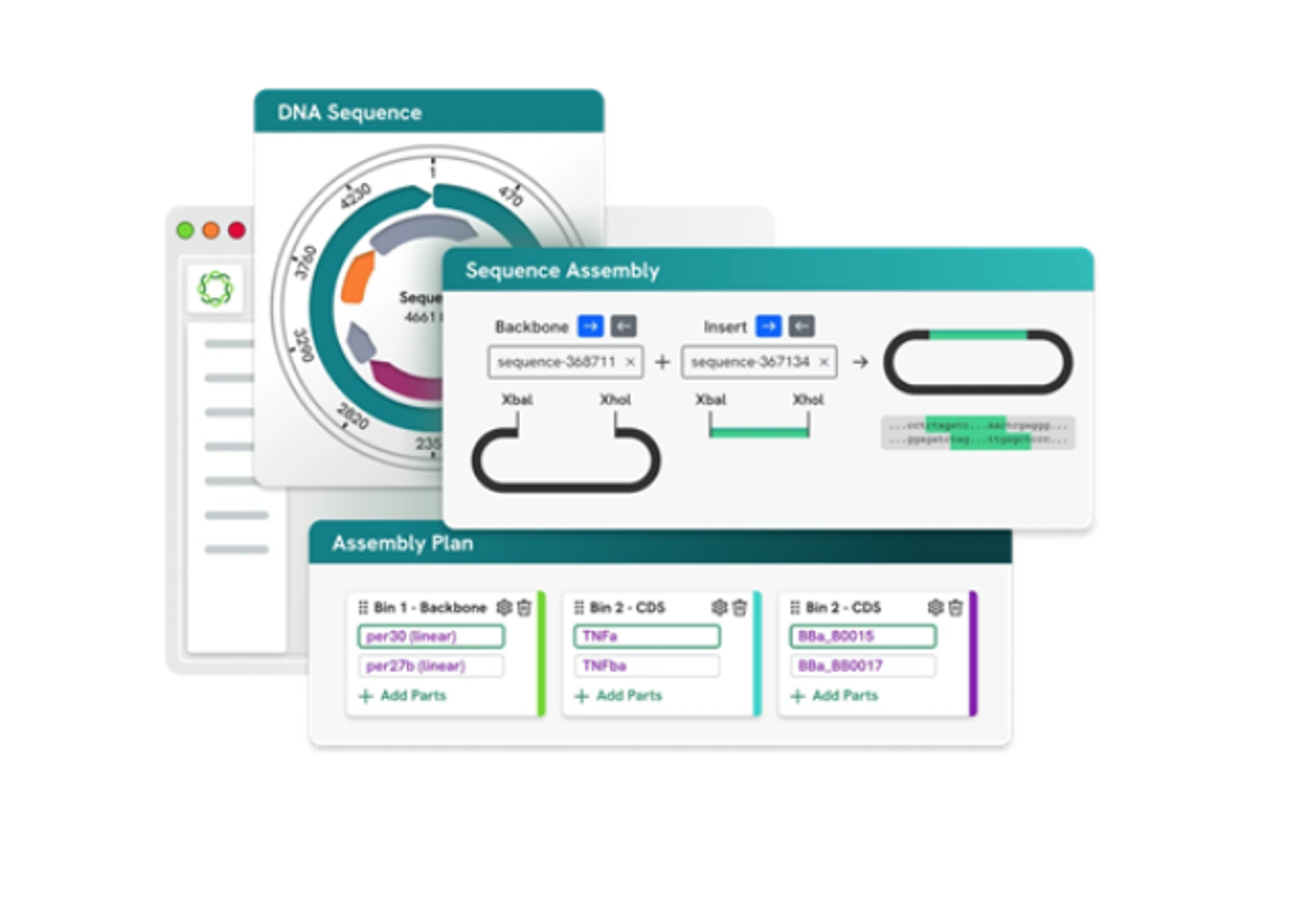
Accel-NGS® 2S MID Indexing Kits (48 reactions)
The Accel-NGS 2S MID Indexing Kits have been designed, optimized, and validated for use withAccel-NGS 2S DNA Library Kits on Illumina platforms, and aid in low frequency variant detection, as well as accurate de-duplication of single read sequencing and sequencing from samples with non-random fragmentation. Get the most out of your NGS sequencing data with this powerful tool. Improve detection of low frequency alleles…

The supplier does not provide quotations for this product through SelectScience. You can search for similar products in our Product Directory.
The Accel-NGS 2S MID Indexing Kits have been designed, optimized, and validated for use withAccel-NGS 2S DNA Library Kits on Illumina platforms, and aid in low frequency variant detection, as well as accurate de-duplication of single read sequencing and sequencing from samples with non-random fragmentation. Get the most out of your NGS sequencing data with this powerful tool.
- Improve detection of low frequency alleles
- Identify unique library molecules
- Retain more sequencing data
- Designed for exome sequencing
- Compatible with circulating, cell-free DNA
Supported DNA Sequencing Applications:
- Whole exome sequencing (WES)
- Deep targeted sequencing
- Cell-free DNA sequencing
- Single-end sequencing, such as ChIP-Seq
Accel-NGS® 2S DNA Library Kits
The application notes provides an overview of Accel-NGS® 2S DNA Library Kits. Accel-NGS 2S DNA Library Kits produce high quality libraries with an all-inclusive, easy-to-use format. Applications of the Library Kits, including use in sequencing liquid biopsy samples and Pico-Scale ChIP-Seq are also covered.
Incorporating Molecular ID Technology with Accel-NGS<sup>®</sup> 2S MID Indexing Kits
This short presentation describes what molecular identifiers (MIDs) and what life science applications can benefit from MIDs. These applications include low frequency variant detection, deduplication from single read sequencing (eg. ChIP-Seq), deduplication from non-random fragmentation (eg. cell-free DNA). It also details the structure of Swift Molecular IDs, describes the bioinformatic analyses that can be carried out with them and gives important guidelines for the kits use.
Accurate De-Duplication of Sequencing Data with New Molecular IDs from Swift Biosciences
Kate Cunningham, PhD, Applications Scientist at Swift Biosciences, introduces the new range of molecular IDs for Accel-NGS® 2S library kits. Hear how the molecular IDs label each individual library molecules, to enable low frequency variant detection, as well as accurate de-duplication of single read sequencing and sequencing from samples with non-random fragmentation. The technology is ideal for applications including whole exome sequencing (WES), deep targeted sequencing, cell-free DNA sequencing and ChIP-Seq.