Webinar Highlights: Automated Low-Volume Liquid Handling for Cost-Effective NGS Library Preparation and Single Cell Genomics
9 Aug 2018
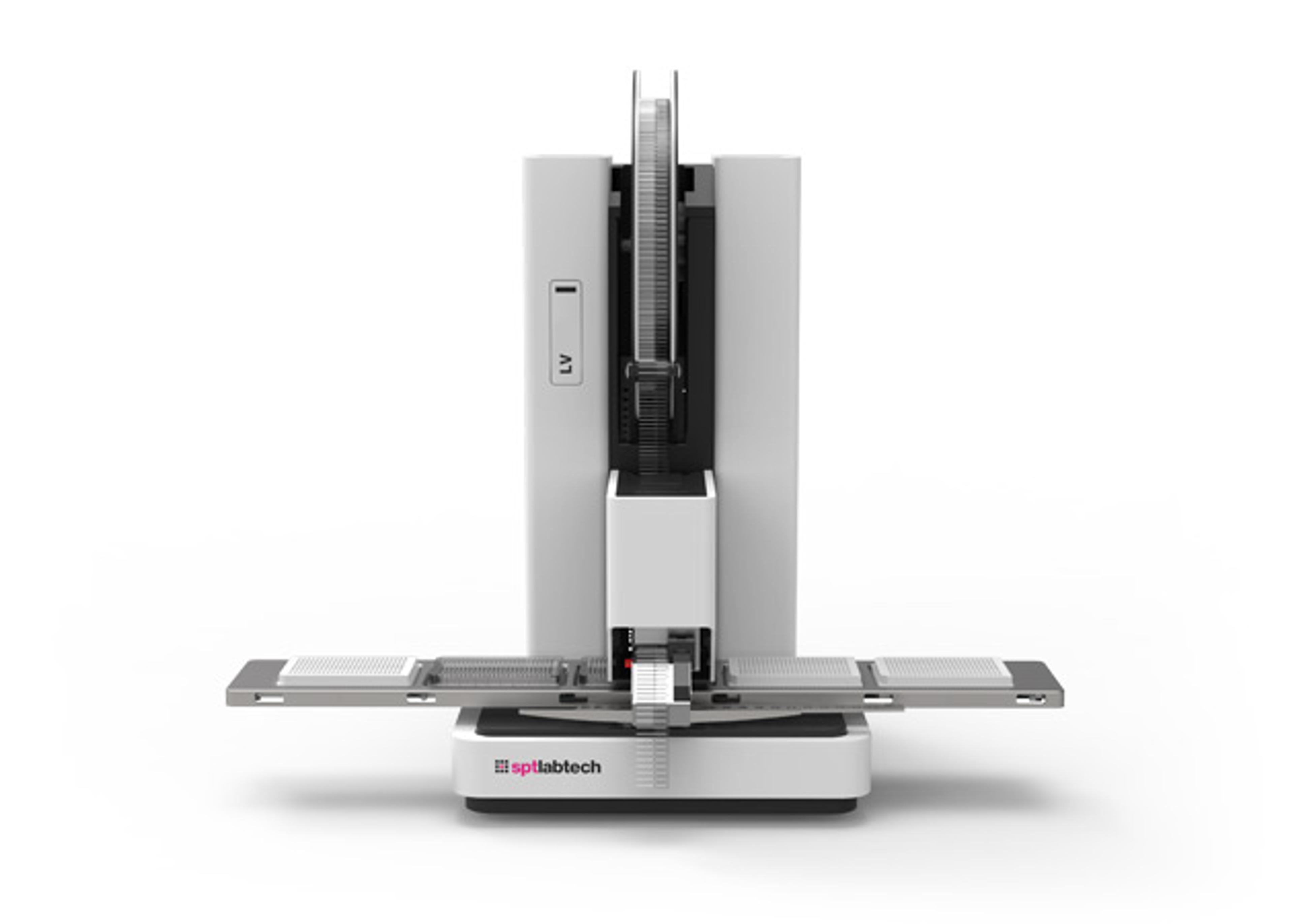
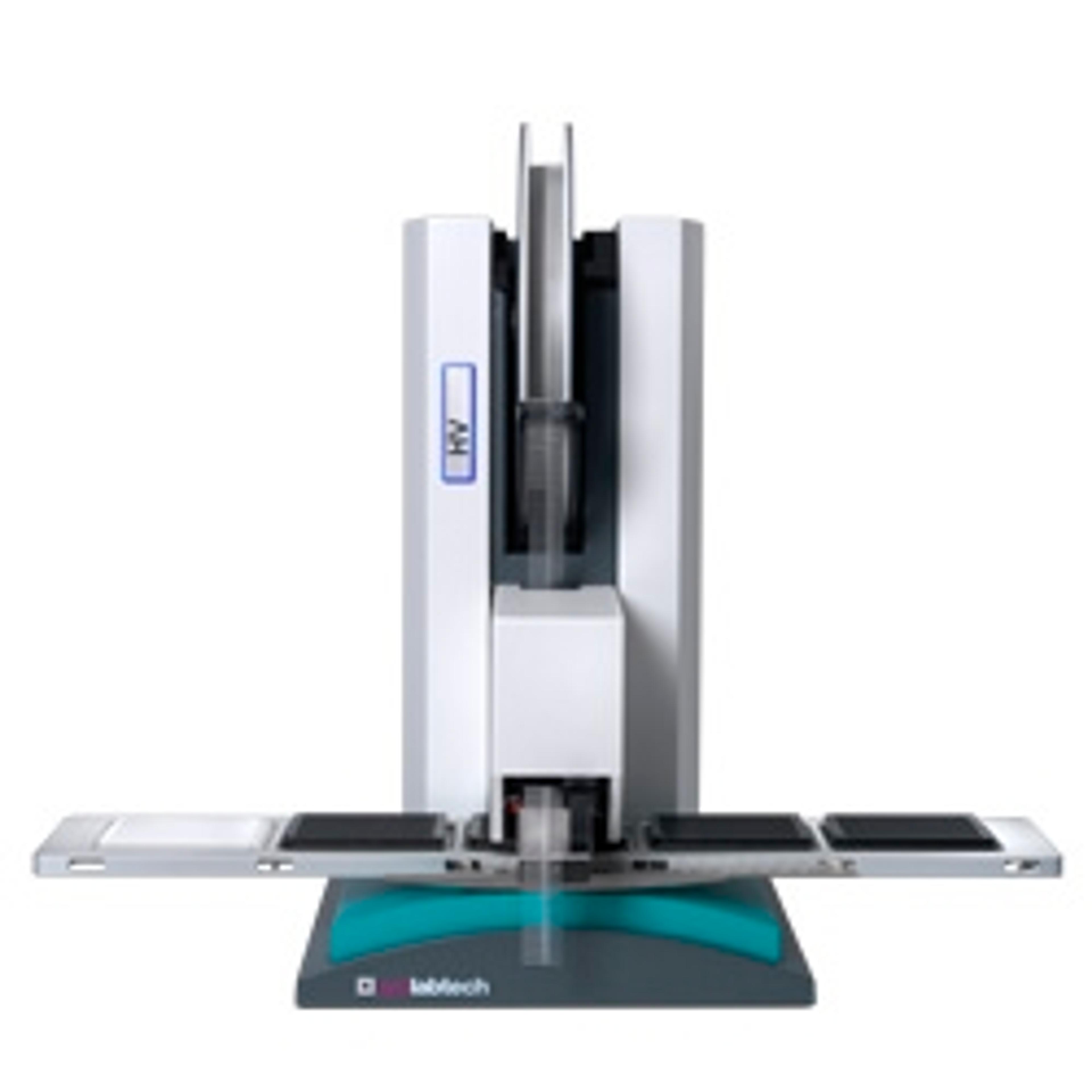
In a recent SelectScience® webinar we were joined for talks by Stuart S. Levine, Director of the MIT BioMicro Center, and Sagar, Postdoctoral Fellow at the Max Planck Institute of Immunobiology and Epigenetics, who discussed how TTP Labtech’s mosquito® liquid handlers are enabling them to perform miniaturized, high-throughput NGS library preparation and single-cell transcriptomics. Now available to watch on demand, this webinar is ideal for both scientists looking to improve the efficiency of their NGS or single-cell analysis pipeline, and those who want to stay up to date with the latest trends in genomics lab automation.
Due to the rapid decrease in the cost of sequencing, sample preparation is often the bottleneck in NGS workflows. Volume miniaturization with automated liquid handlers can potentially solve throughput and cost challenges. However, suitable liquid handlers must provide high accuracy at low transfer volumes, while avoiding cross-contamination. Unlike traditional liquid handlers, TTP Labtech’s positive displacement pipetting technology is highly accurate in the low-microliter to nanoliter volume range, even at high viscosity.
You can catch the webinar on demand here or read on to discover highlights from the webinar’s Q&A session, where we were also joined by liquid handing technology expert, Klaus Hentrich, Global Product Manager Genomics, at TTP Labtech.
Question and Answer Session Highlights
What is the effect of miniaturization on overall library prep costs?
SL: We see reductions of multiple folds, but when you do one twelfth reaction volume, it isn’t one twelfth of the cost. With everything put together, we can typically reduce costs of library prep, etc., at least four-fold if not more.
Sagar: It’s the same for us. When we were doing single-cell sequencing experiments using the original CEL-seq2 protocol, the costs per cell were approximately 10-12 euros per cell. We were able to achieve a six-fold cost reduction to around two euros per cell, meaning we are now able to sequence really large numbers of cells. Since the single-cell sequencing field is really moving forward rapidly, and people are sequencing hundreds or thousands of cells at a cheaper rate, this reduction has become necessary. This reduction in library prep charges has been a major advantage for us.
Which instrument model were you using and were you concerned about evaporation?
Sagar: We are using a mosquito HTS, so the volume range is 25 nL – 1.2 µL. Initially we were worried about evaporation, but we were not really using mineral oil, which did work OK but only because we were using deep-welled 384-well plates and keeping samples on ice most of the time. The problem comes when you have an inactivation step of 70 degrees, which can cause you to lose your droplet – that’s why we switched over to the mineral oil. We prepare the mineral oil plate first with 1.2 µL of mineral oil then dispense 240 nL of the lysis buffer.
I would really recommend if possible that you use mineral oil or Qiagen Vapor-Lock, which is like mineral oil with a water-like consistency. During sorting, the cell will most likely be in the mineral oil, but after centrifugation, the cell will definitely go in the lysis buffer.
What were the major challenges to solve in your workflow?
Sagar: We had the mineral oil on the top so we had to really do a lot of color experiments using mosquito so that we know that the droplet is maintained at the bottom of the well. We optimized our whole protocol using these color-based experiments. The other challenge was the fluorescence activated cell sorting (FACS), where we have to make sure that the machine is calibrated and able to sort each single cell at the center of the well.
SL: It really varies with the chemistry. Some of the chemistries miniaturized very easily and work very quickly, while other kits just haven't worked. Some of the RNA-seq chemistries we've been starting with had significant primer-dimer problems that occur only when we're miniaturizing and the better solution to that, for us, is finding the right chemistry.
What is the dropout or doublet rate in your workflow?
Sagar: For the doublet rate, because we are using FACS to sort cells via forward- and side-scatter, we can exclude doublets almost completely, which is not possible if you use a droplet-based technique where the doublet rate is high. The dropout rate depends on various factors, for example there are some cell types where we can retain approximately 90% to 95% of the cells, but with some larger cells such as hepatocytes or cardiomyocytes we have a poor recovery. This is most likely due to the cell size where the FACS nozzle becomes a critical factor, so the equipment is also important.
SL: For dropouts, sometimes we will get samples that fail, but that's not a noticeably different rate. In fact, it's probably lower on the robots than by hand.
What's the difference between the mosquito and other liquid handlers?
KH: The key difference is the accuracy of the low volumes. Sagar’s system works from 25 nL – 1.2 µL and Stuart's system between 0.5-5 µL. The positive displacement pipetting without any air gaps makes it possible to achieve these high accuracies and handle viscous liquids at small volumes. You can even dispense the Vapor-Lock, mineral oil, and high-glycerol enzyme mixes without any liquid classification. Also ease-of-use, robustness, and the disposable pipette tips that can be automatically changed to avoid cross contamination.
Sagar: When you work with single cell RNA-seq where you have different barcodes, you want an RNase-free environment and also avoid barcode contamination. So it was really important for us to have disposable tips like mosquito has. And it’s very fast compared to other liquid handlers we have seen. The speed was also important to us when we bought the machine.
Register to watch the full webinar on demand or learn more about the mosquito liquid handlers from TTP Labtech.


