Biomarker detection in 3D cell models using automated immunofluorescence staining
Watch this webinar on demand to learn how to automate your immunofluorescence staining workflows
10 Apr 2022
Expert insights

Physiologically relevant 3D cell models are essential for drug discovery and preclinical research due to their functional and architectural similarity to solid tumors. One of the challenges faced by researchers is that many of the assays using these precious samples tend to be manual and tedious. Using proprietary microfluidics technology, Protein Fluidics has created the Pu·MA System for automated complex 3D cell-based assays.
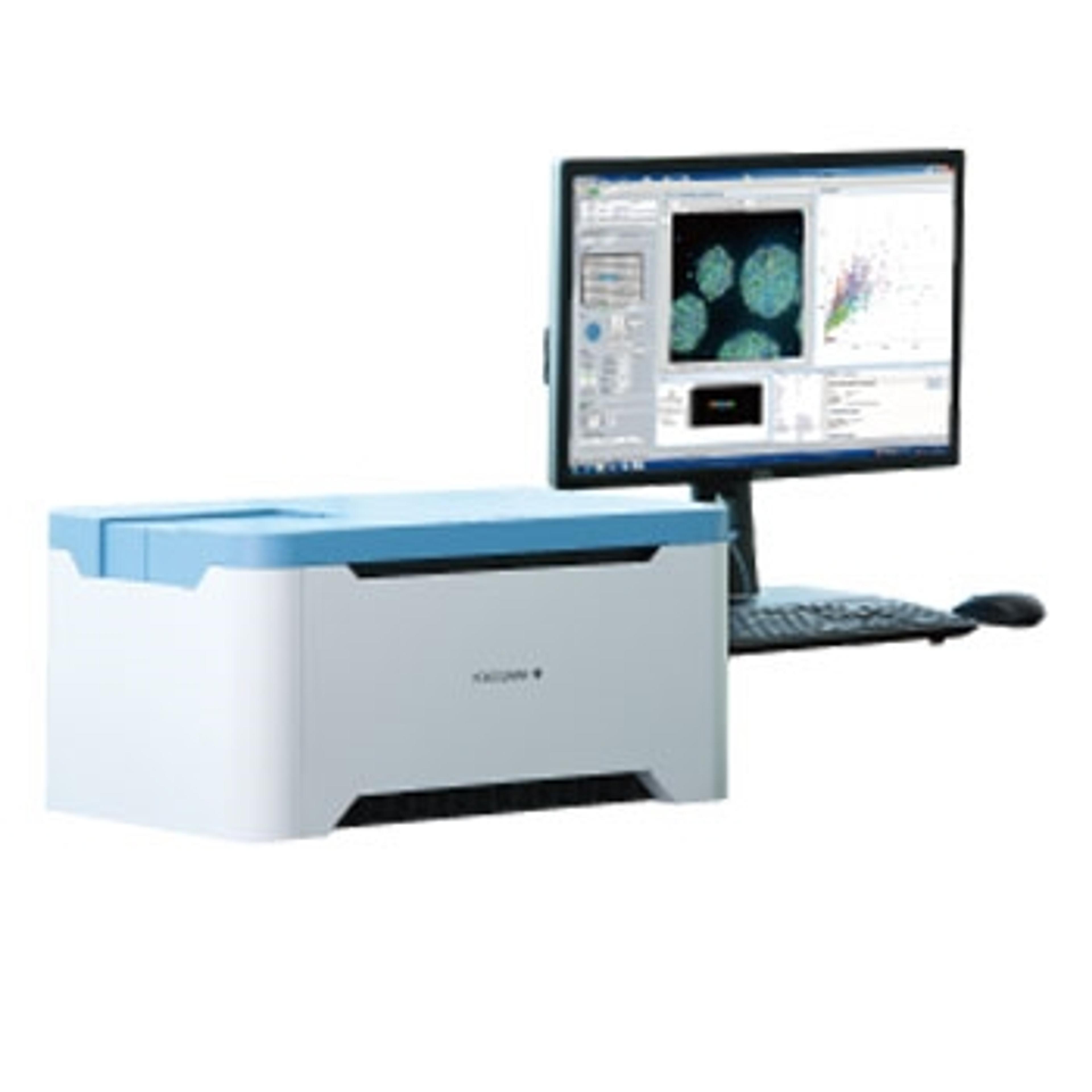
In this SelectScience® webinar, now available on demand, application scientist Dr. Katya Nikolov presents her work on combining this novel automation technology with Yokogawa’s high-content imaging systems for biomarker detection in 3D cell models. Nikolov demonstrates the utility of an automated immunofluorescence (IF) staining workflow followed by confocal imaging within the Pu·MA System flowchips. This automated workflow enables quantitative assessment of biomarkers which provides valuable data for further understanding disease mechanisms, preclinical drug efficacy studies, and for use in personalized medicine.
Watch on demandRegister now to watch the webinar on demand, and read on for highlights from the live Q&A session:
What fluorophores did you use in your assay?
KN: It can be different sets of fluorophores depending on the confocal system or the microscope that you're using and depending on the capabilities of the system.
For example, we use Hoechst for identification of the nuclei. We also use different sets of primary antibodies conjugated with different kinds of Alexa fluorophores, such as Alexa 488 conjugate. We also use LipidTOX stain.
Are the plates also compatible with confocal microscopy or an Operetta system?
KN: The frame and flowchips have the same dimensions as standard three to four-well plates or just standard cell culture plates. You can put it in any type of microscope, or height-intense scanner, or in a plate reader. So, any instrument that you normally would use with standard culture plates.
What type of fixative did you use for the Pu·MA System? And did you experience any challenges with Matrigel or VitroGel?
KN: In terms of fixative, we use standard protocol. In most of the cases, we use 4% paraformaldehyde (PFA). In terms of Matrigel and VitroGel, the way we handle and load samples in these gels did require certain optimization. But we optimize protocols so that there is no interference with the microfluidic part of the flowchip. So, I would say no.
The protocol was optimized so that we were able to obtain pretty good images with high quality, and at the same time maintain samples within the protected chamber surrounded by matrix. Depending on the sample, it might require some optimization, but we don't have any issues with that.
Is it possible to create organoids from primary cultures also derived from patients, or does the culturing of patient cells not allow keeping the heterogeneity of tumor cells? If so, can you keep the organoids to later transplant into mice, for example?
KN: Yes, it is possible to create organoids from primary cells. There are different protocols and approaches that scientists tend to use. However, the current design of the flowchip is not designed for creating and culturing organoids, so they're usually created elsewhere and then we transfer them to the flowchip for the assays. Normally we wouldn't take organoids out; it may be possible, but it's not the goal of the system. We just create a set of assays, and we wouldn't take them out for the flowchip to inject into animals.
Usually in IF, we need to wash after the first antibody several times. Does the Pu·MA System allow for multiple wash steps?
KN: Yes. The protocol is developed in such a way that we do all the standard steps of IF staining. We start with blocking, then incubation with primer. You can adjust the time of this incubation. Then we typically have three washes, and then there's a step to do secondary antibody incubation, and washes after that. So, you have the capability to manipulate the number of washes, but they are all incorporated in the protocol.
Do you have any experience related to other 3D culture models of steroids such as embedding the cells in Type I collagen in Pu·Ma’s chambers?
KN: We have worked with different hydrogels, engineered hydrogels, and mostly Matrigel. Protocols in the Pu·MA System can be optimized for different types of matrices as long as it doesn't interfere with the microfluidics. So generally, this system is matrix-agnostic, but we haven't tried all of those that are available yet.
To learn more about the Pu·MA System, watch the webinar on demand>>
SelectScience runs 10+ webinars a month, discover more of our upcoming webinars>>