Collaboration and CRISPR tools move scientists closer to a cure for sight loss
See how researchers are uncovering the mechanisms of retinal degeneration and identifying new targets for therapeutic intervention
6 Mar 2023
Editorial article

Approximately 2.2 billion people worldwide suffer from vision impairment, with Age-related Macular Degeneration (AMD) - caused by the death of photoreceptors cells primarily in the macular region of the retina - being the leading cause of blindness among elderly individuals. Genome-wide association studies (GWAS) have exposed the involvement of numerous common genetic variants in AMD. However, efforts to understand the underlying mechanisms of these variants have been thwarted by the lack of a suitable disease model that accurately mimics the right disease context.
Organoids, the self-organizing three-dimensional cellular models derived from stem cells, have opened the possibility of replicating the human eye to closely examine and study vision loss. Now, these models are being paired with the latest CRISPR technologies to introduce precise genetic edits and examine the role of individual variants on the mechanisms of retinal neurodegeneration.
In this SelectScience® interview, we speak with Dr. Yu Holly Chen, Assistant Professor at the University of Alabama at Birmingham, and Dr. Rinki Ratnapriya, Assistant Professor at the Baylor College of Medicine, to learn how they are combining their respective expertise in developing in vivo models for retinal diseases and employing sequencing-based genome-wide methods to advance our knowledge of AMD disease mechanisms and explore potential therapeutic approaches.
Closing in on genetic culprits
Genome-wide association studies have identified 34 loci associated with AMD. However, as most of these variants reside in the non-coding region of the genome, identifying causal variants, target genes, and the mechanisms by which they mediate disease phenotypes has remained a key challenge.
Retinal organoids, which broadly mimic the development and morphology of the human retina, offer a means to recapitulate disease mechanisms and progression in vitro and to interrogate the role of individual AMD-associated variants. “As humans harbor many different variants, studying these in lab-grown cells offers a unique window into studying the disease mechanisms,” explains Ratnapriya. “This led to our collaboration with Dr. Chen, who is an expert in developing in vivo models for retinal diseases.”
Chen’s research focuses on inherited retinal degeneration, and specifically, how genetic mutations lead to defects in sensory structures – known as primary cilia – within the eye. “We use retinal organoids derived from human pluripotent stem cells, which form miniature 3D tissues that structurally and somewhat functionally resemble in vivo retina,” she says. “These offer a promising platform to study disease mechanisms and develop therapies in a human context.”
“Rinki and I both had our postdoctoral training in the same lab at the National Eye Institute (NEI), and our expertise and research interests are complementary to each other,” Chen continues. “By working together, we hope to couple this human pluripotent stem cell-based platform with genomic techniques to advance our knowledge of AMD and explore various therapeutic approaches.”
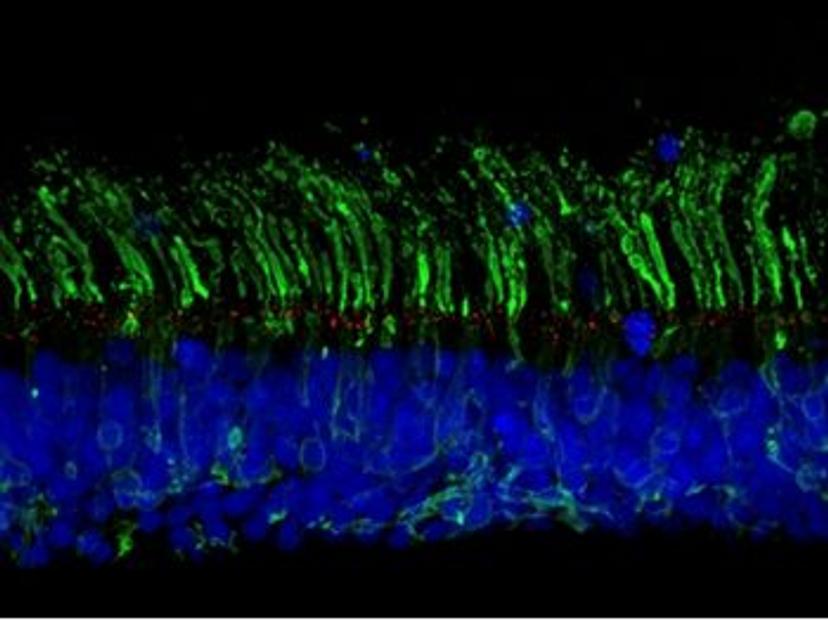
CRISPR-engineering of isogenic organoids
Chen and Ratnapriya’s collaboration aims to uncover the mechanisms by which non-coding variants regulate the expression of genes that may play a role in AMD. To this end, the researchers intend to generate isogenic organoid models – cell lines with the same genetic makeup, bar specific genetic modifications of interest. “We plan to generate isogenic lines with AMD-associated variants using CRISPR gene editing”, says Ratnapriya. “In doing so, we hope to study the long-term impacts of these variants on gene expression, regulatory networks, and disease mechanisms.”
Ratnapriya goes on to explain the value of CRISPR technologies in this work. “CRISPR-based gene editing, as well as gene activation (CRISPRa) and gene inhibition (CRISPRi), offers a host of strategies to study regulatory elements harboring causal variants in disease-relevant cell lines to investigate their functions, target genes and overall role in causing disease phenotypes,” she says.
“Compared to other genome-editing tools such as TALEN, CRISPR is a more versatile and easier-to-design approach,” adds Chen, noting how advances in CRISPR technologies have greatly simplified the generation of isogenic in vitro models of human disease.
“Besides my collaboration with Rinki, I’m also using CRISPR tools to generate isogenic and conditional knockout hPSC-derived models to dissect the role of specific genes and cell types in disease progression,” she adds.
Experimental challenges
Despite advances in the precision and efficiency of CRISPR-based gene editing, Chen and Ratnapriya’s research is not without its challenges. “Like most complex diseases, AMD is also caused by the cumulative impact of multiple causal variants, which remain technically challenging to model in cell culture,” says Ratnapriya. “Using CRISPR to follow up candidate genes also requires previous knowledge regarding which functional assays are the most disease-relevant, which are not always very well understood.”
“Some researchers hesitate to use CRISPR-based gene editing tools due to their concern about off-target effects or other pitfalls,” adds Chen. “While I understand those are the current shortcomings of the approach, it's important to remember that every new technology has initial difficulties, and these can be addressed and improved over time. For example, now different strains of Cas9 proteins have been developed to reduce off-target effects and improve efficiency. If we didn’t try anything new, there would be no improvement.”
The future of gene editing
Beyond developing disease models, Chen and Ratnapriya see great promise in the use of CRISPR Cas9-mediated genome editing in therapeutic applications, where it could potentially be used to correct disease-causing mutations associated with sight loss in humans. “With improvements in efficiency and specificity, in the future, we could directly inject the gene-editing machinery into disease-relevant tissues to correct the mutations on site,” Chen enthuses.
“There are several challenges associated with gene editing technology applications in humans, such as off-targeting, polymorphism, delivery method optimization, and ethics. However, technical advancement in gene editing presents a real opportunity to prevent, treat, and cure inherited diseases,” adds Ratnapriya.
“Gene editing has expedited the research in gene expression regulation, epigenetic modification, therapeutic drug development, functional gene screening, and gene diagnosis,” she continues. “The long-term goal of utilizing genome editing in disease prevention and treatment remains an attractive avenue, and my advice to other researchers is to keep pushing the envelope and ask pertinent questions to make gene editing available as a next-generation therapeutic option.”
Organoids and CRISPR gene editing: Tools for success
When conducting organoid- and CRISPR Cas9-based research, the use of high-quality reagents is imperative. The organoid models Chen is developing are grown using culture media, growth factors, and reagents from Thermo Fisher Scientific. Find out more about their solutions at thermofisher.com/geneediting
Learn more about how CRISPR gene editing technology is supporting researchers in their work:
- Dr. Sergio Casas-Tintó, head of the Drosophila Models of Human Disease Unit at the Institute for Rare Diseases Research of ISCIII, is combining CRISPR gene editing with alternative animal models to uncover the genes responsible for rare and ultra-rare diseases.
- Dr. Krishanu Saha, associate professor of biomedical engineering at the University of Wisconsin-Madison, highlights the potential of CRISPR to improve CAR T-cell efficacy for solid tumor treatment.