Integrative Microscopy Approaches for Enhanced High Resolution Imaging
Discover the advances in imaging technology that are accelerating plant biology
15 May 2016

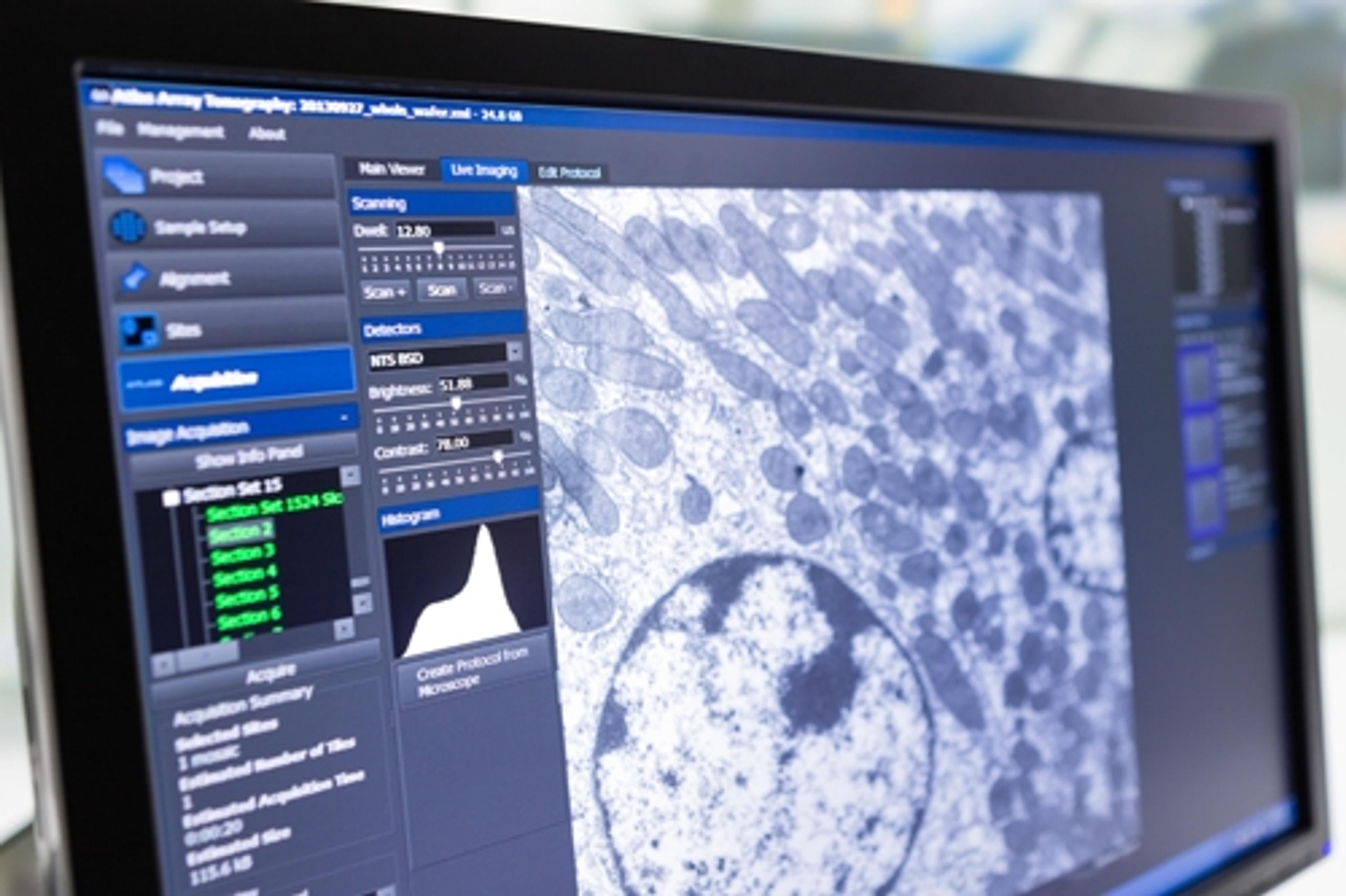
Image: ZEISS Microscopy
Professor Chris Hawes, Professor of Plant Cell Biology, Research Lead and Head of Doctoral Training Programme at Oxford Brookes University, told SelectScience® about his plant cell imaging research. Prof Hawes explained how advances in imaging technology, such as correlative microscopy, are enabling the elucidation of plant structure from tissue level down to subcellular organization.
Prof Hawes’ research group is interested in the structure and function of plant cells, with a focus on secretory pathways, the endoplasmic reticulum (ER) and the Golgi apparatus. One of the group’s current projects is investigating the subcellular localization of auxin production, a hormone with a key role in plant development. Dr Verena Kriechbaumer and Prof Hawes predict that auxin is made on the ER membrane surface, rather than free in the cytosol, which is a new concept in plant biology with potential implications for food production and pesticides.
High Resolution Imaging
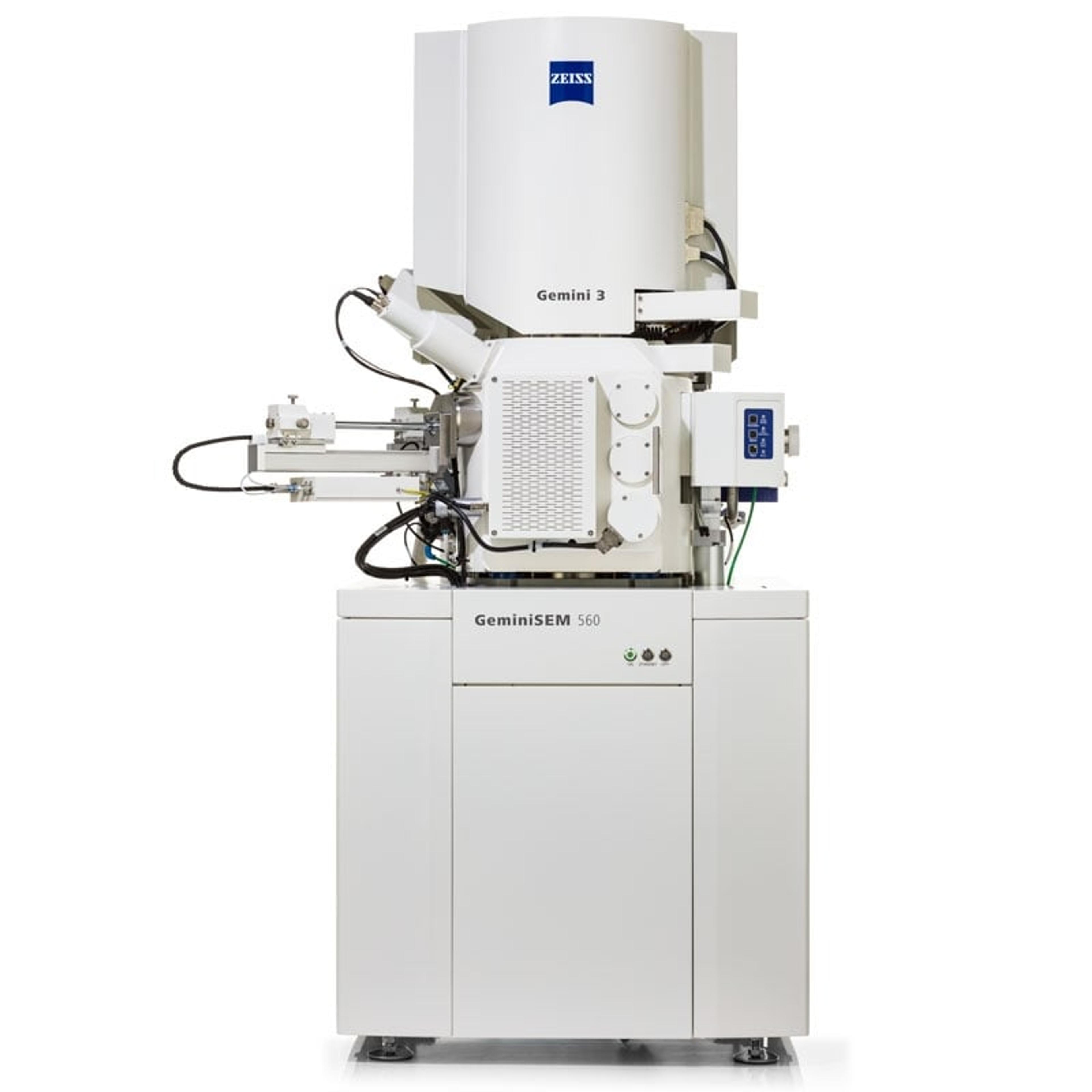
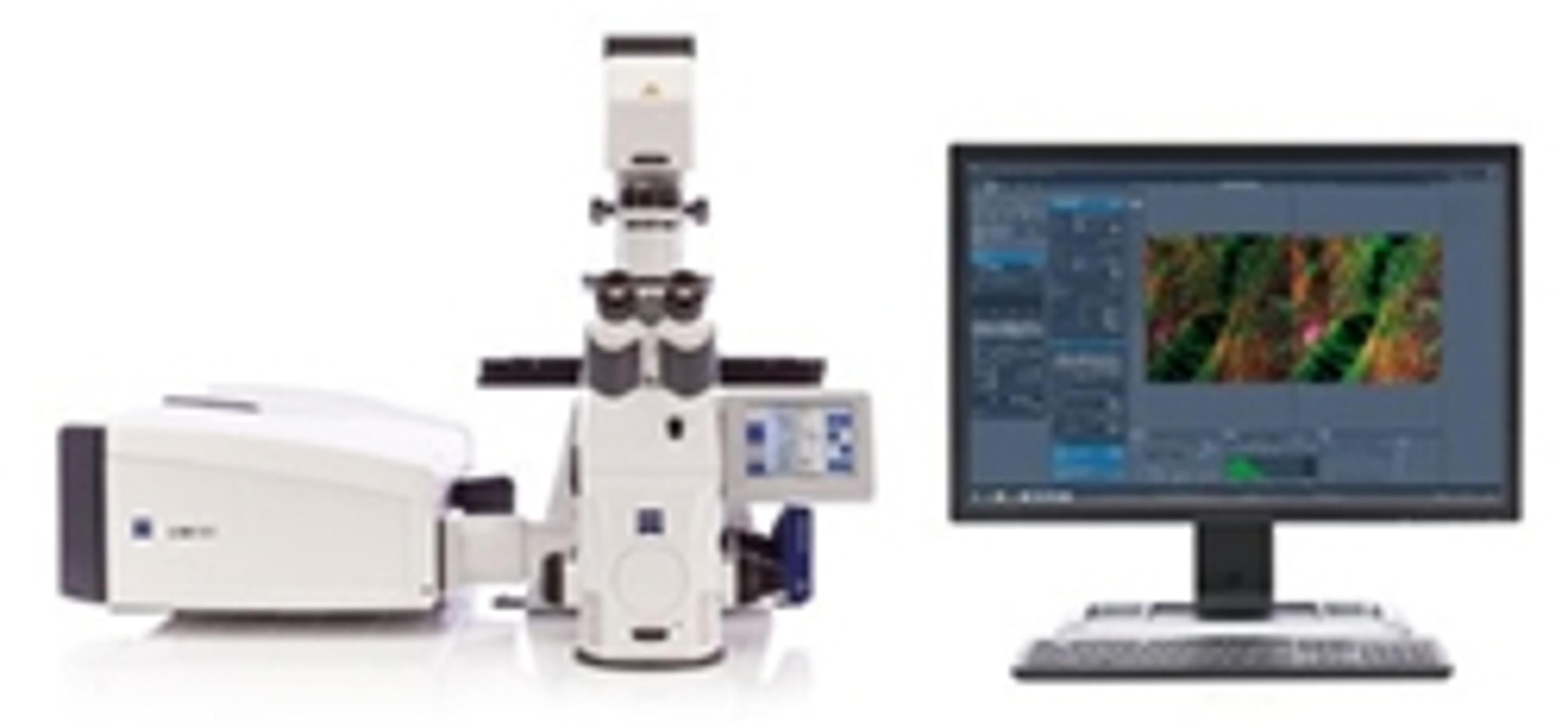
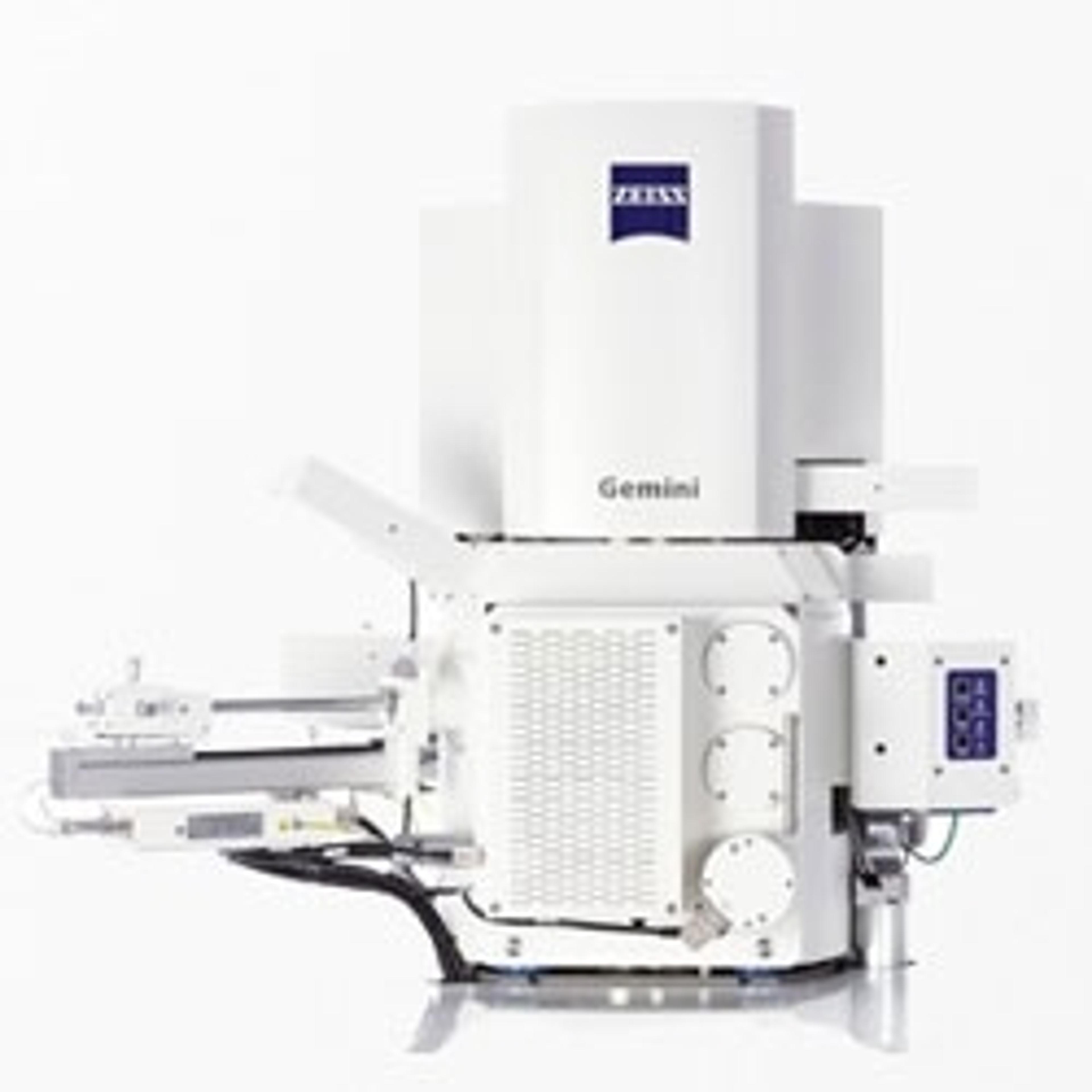
Kriechbaumer and Hawes are currently using two strategies to study clustering of auxin associated metabolites on the ER membrane network. Fluorescently labelled proteins are imaged using ZEISS LSM 880 with Airyscan, in live cell confocal microscopy, followed by serial block-face scanning electron microscopy with ZEISS Merlin 3View FE-SEM. Superimposition of these images enables the subcellular location of the tagged proteins to be estimated. The group are also using Total Internal Reflection Fluorescence microscopy (TIRF) to image membrane surface proteins, at the single molecular level and in the future will label proteins in metabolic pathways with differently colored fluorescent labels, which will allow clustering of individual molecules to be detected. However, as Prof Hawes explains, “You can look at hundreds of specimens by fluorescence microscopy on the confocal and you can look at hundreds of specimens on the electron microscope, and you can get a pretty good idea how to correlate the light with the electron microscopy but you can’t say this is in the very same cell”.
Advantages of Correlative Microscopy
Correlative microscopy enables a greater level of precision. Light and electron microscopy are integrated using specialized software that automatically superimposes the light and electron images, so that the exact same location is being imaged. This has important implications where analyses need to be localized to targeted cell interactions, to avoid the detection of neighboring cell effects. Pending funding, Prof Hawes hopes to switch to a correlative workflow to enhance his plant imaging studies. Prof Hawes also believes that the recent development of fixation resistant fluorescent proteins is a major breakthrough that will greatly enhance the flexibility of high resolution correlative imaging, by enabling fluorescently labelled fixed tissue to be imaged using confocal microscopy, prior to electron microscopy.
Future Plans
Prof Hawes future goals involve making reconstructions of the 3D organization of plant tissue using 3D scanning electron microscopy. He aims to reconstruct the endomembrane system, which could have major implications for protein production and crop productivity. As Prof Hawes explains, higher resolution techniques allow the study of the organization in living cells to be taken to the next level, providing better information on where individual proteins are located and how they interact.
Find out more about this research on Prof Hawes' website and visit our Correlative Microscopy and 3D microscopy special feature pages for the latest developments.