KAPA Library Quantification Kits
Kapa Biosystems, Inc.KAPA Library Quantification Kits contain all the reagents needed for the accurate, reliable and reproducible qPCR-based quantification of NGS libraries prior to pooling for capture or flow cell amplification.
Kits contain KAPA SYBR® FAST qPCR Master Mix, optimized for high-performance SYBR Green I-based qPCR. The engineered KAPA SYBR FAST DNA Polymerase contained in the Master Mix amplifies GC- and AT-rich DNA fragments of different lengths with similar efficiency, making it the only qPCR Master Mix capable of accurate qPCR-based quantification of NGS libraries. The pre-diluted set of DNA Standards contained in each kit is carefully quality-controlled to ensure lot-to-lot consistency and avoid data drift over time. Optimized Primer Premixes ensure optimal PCR efficiency.
Features:
Accurately quantify PCR-competent sequencing templates
qPCR only “counts” those library molecules that can be sequenced
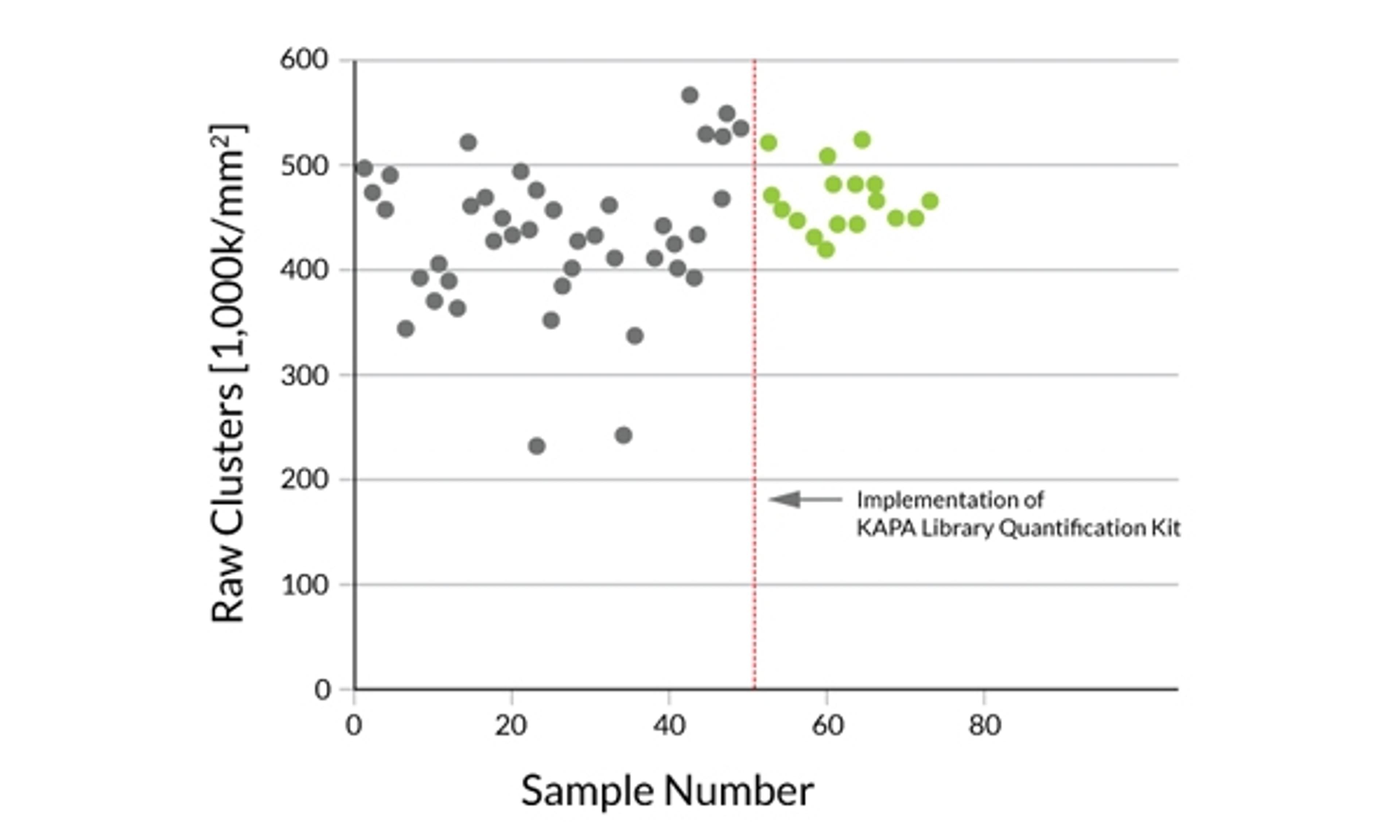
More consistent cluster densities or template-to-bead ratios enable optimal utilization of sequencing resources
Improve pooling for multiplexed sequencing
Library concentrations determined by qPCR are more reliable compared to other methods
The engineered KAPA SYBR FAST qPCR Master Mix enables accurate pooling of libraries with extreme GC-contents or longer inserts
Eliminate drift with carefully quality-controlled DNA Standards
Prediluted DNA Standards eliminate quantification errors associated with preparation of standard curves using in-house standards
KAPA DNA Standards undergo strict quality control to ensure lot-to-lot consistency and eliminate data drift over time
Improve throughput with automation
Library quantification assay is compatible with 96- and 384-well format
Library dilution, reaction setup and data analysis can be automated for HTP pipelines
Applications:
Quantification of libraries prior to pooling for multiplexed sequencing
Quantification of individual libraries or library pools prior to cluster amplification or emPCR
Quantification of libraries prior to and after hybridization capture
Quality control, optimization and troubleshooting of library construction processes or workflows