Ultra-Sensitive Digital Method Detects Rarest of Genome Mutations
Dr. Bruce Conklin, UCSF, discusses the potential of genome engineering as a therapy for genetic diseases
4 Dec 2017

Dr. Bruce R. Conklin, M.D., is an Investigator at the Gladstone Institutes and a Professor in the UCSF Division of Genomic Medicine. His lab uses induced pluripotent stem cells (iPSCs) to study the genetics of cardiovascular disorders.
In this SelectScience® interview, we speak with Dr. Bruce Conklin, Gladstone Investigator, and Professor in the Division of Genomic Medicine at University of California, San Francisco, whose research was instrumental in developing a robust, sensitive method to detect edited genes within thousands of copies of genomes, and propelled the field of genome engineering in iPSCs.
The Conklin lab uses human iPSCs to model genetic diseases. “There are two models of iPSCs in translational research,” says Conklin, “one is where we start off with a healthy line and impart mutations on that background.” This ‘healthy’ model is analogous to an inbred wild-type BL6 mouse on which mutations are imparted to create a disease phenotype. In such models, it is easier to compare mutations because they all have the same genetic background. “The second iPSC model is obtained from patients and represents a human disease.” Correcting patient-derived iPSCs is much more difficult as every cell line generated is different. These diverse genetic backgrounds make it challenging to determine the molecular basis of the allelic series i.e. the mutant alleles of the gene. However, if the purpose of the study is to actually treat the patient from whom the iPSCs are derived, it is valuable to use the patient’s own cells as controls. “We use either of these models depending on what the genetic problem is – each has its own place in research,” says Conklin.
When Conklin began work with iPSCs in 2008, he soon realized that genome engineering was imperative to advancing his research. “When we started working with patient-derived iPSCs, there were inadequate controls that were genetically identical to one another, sparing the gene that was mutated. So, we began using genome editing for creating better disease models, particularly for cardiac disease, by making larger allelic series of point mutations in iPSCs with a common genetic background,” Conklin explains. His efforts have resulted in a wild-type iPS cell line called WTC which is used widely throughout the world as a control. For instance, the Allen Institute uses WTC as the background for a large series of GFP-tagged genes.
“Genome editing was extremely powerful in the sense that when we started working with iPS cells, there were only eight targeted lines in human embryonic stem cells. Now, we have well over a 100! Without gene editing, this development wouldn’t have been possible."
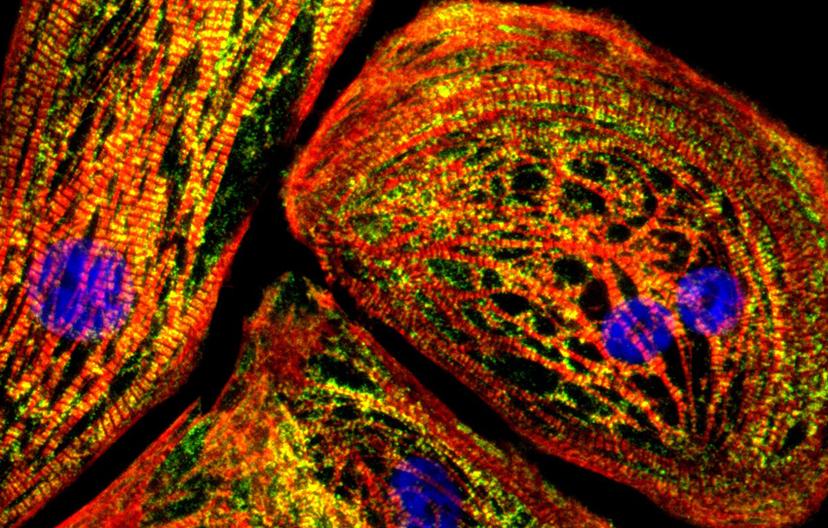
Human iPSCs were differentiated into cardiomyocytes. Red: anti-TNNT2 (the TNNT2 gene codes for cardiac troponin T, found solely in cardiac muscle). Green: anti-BAG3 (the BAG3 gene is implicated in dilated cardiomyopathy). Image taken by Dr. Luke Judge, M.D. Ph.D. of the Conklin Lab
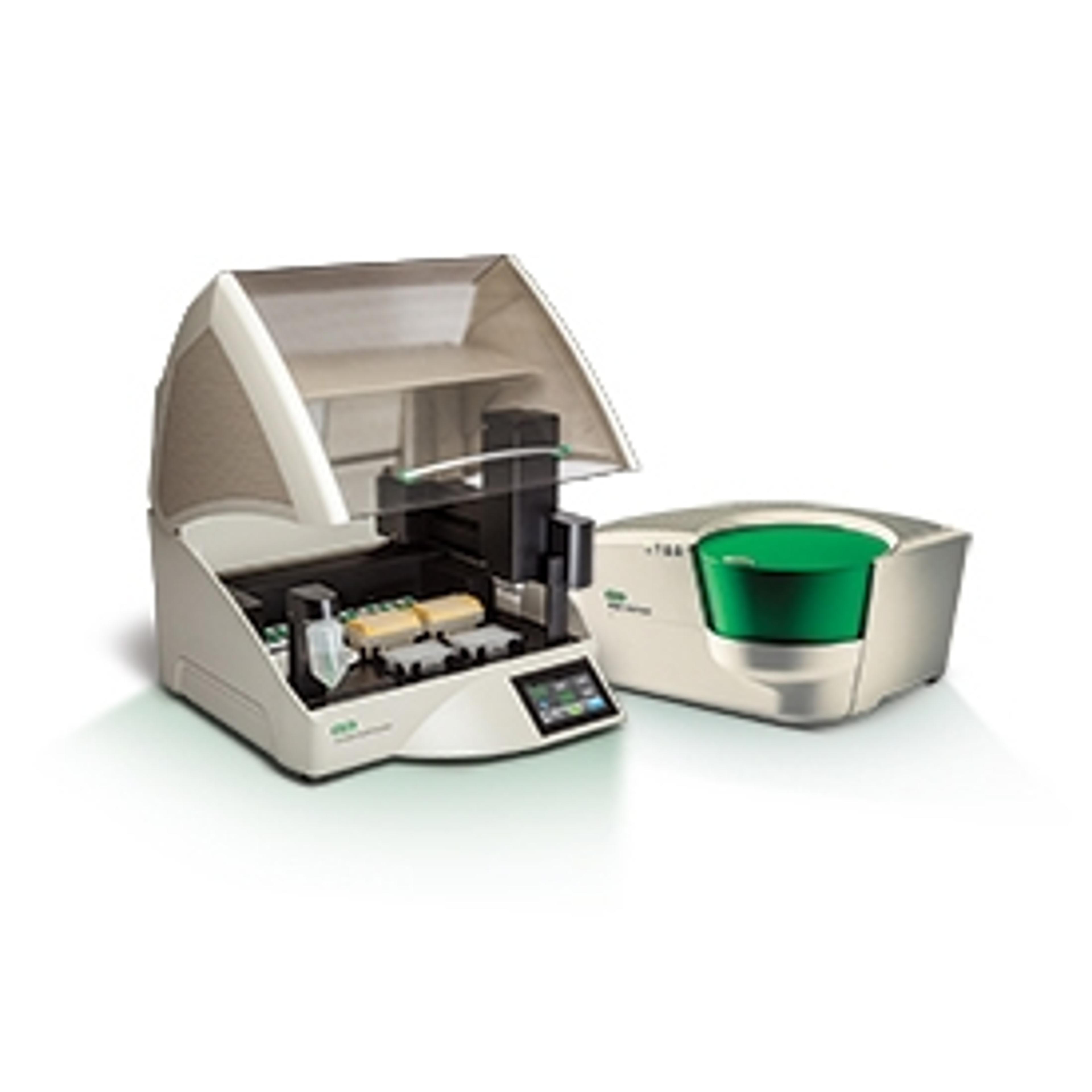
Genome editing methods, however, are only as powerful as the protocols devised to detect and quantify them. With the advent of droplet digital PCR (ddPCR) — a method where each PCR sample is emulsified with oil and partitioned into 20,000 nanoliter-scale droplets so as to detect even the rarest of genetic events — Conklin’s lab developed a robust, sensitive protocol to detect single nucleotide substitutions, published in Nature Methods, 2014 and Cold Spring Harb Protoc, 2016.
“We started using ddPCR back when we were doing single base changes in DNA. It’s incredibly powerful – for each droplet, you’re measuring a single copy of DNA,” says Conklin. “For instance, it is able to reach into 2,000 genomes to detect a rare single base pair change in one of the alleles.”
Compared to using a Southern blot or qPCR, Conklin further comments on his choice for ddPCR: “There are other systems that are called digital, but this is really the only system I know of where you’re detecting a single molecule of DNA. You either detect it or you don’t. Some of the other methods are actually population-based i.e. they detect relative abundance of thousands of molecules. With the ddPCR, we are able to detect both the copy number variation and single point mutation, which is very valuable.”
Shared among 30 different researchers in the Gladstone Institute, the Bio-Rad ddPCR QX100 instrument stays active. “There are some machines that are difficult to maintain in a group setting, but not the QX100 – this one is pretty robust. The method is not complicated, and the software is pretty intuitive,” says Conklin.
As researchers in the Conklin lab isolate clones of iPSCs to model a disease, they perform ddPCR to detect the copy number of the gene of interest using fluorescent tags. If one cell line has one copy, and another has five, this feedback is then used to optimize transfection protocols on the latter to generate a single-copy cell line. “As we’re trying to understand how one gene insertion cassette works with different iPSC patient lines, it’s very difficult to make a comparison when one cell line has five copies and the other has one. We need to have the same number of copies per cell, and ddPCR is the fastest way to measure that as it can be done in 96-well plates,” Conklin explains.
The Conklin lab has ongoing projects to model cardiac, neural and retinal diseases in vitro using iPSCs, an indispensable step for precision medicine. “Each of our genomes is intrinsically very different,” Conklin says, looking ahead to the future of genome engineering. “I call it ‘genome surgery’. Just as a surgeon would approach each patient differently for a procedure such as a heart transplant, that is how we need to think about therapeutic genome-editing to target diseases.”
References:
Miyaoka Y et al. (2014) Isolation of single-base genome-edited human iPS cells without antibiotic selection. Nat. Methods 11:291.
Do you use ddPCR in your lab? We'd love to hear from you. Share your expertise and leave an opinion here.

