Top tips for diagnosing PCR issues
What can go wrong in the lab when the science and theory are correct?
11 May 2022

Polymerase chain reaction (PCR) is a revolutionary method of DNA amplification applied to a wide range of genetics-based scientific investigations, but when performing this form of highly sensitive microscopic amplification, it is vitally important that all aspects of the workflow are free from error. Any imperfection in the compounds or oligonucleotides involved, or contamination of the sample material, can be amplified a billion times over along with the production of new DNA copies, and so it is important to take the right preventative measures to ensure results are accurate, reliable, and reproducible.
John Winsor is a Product Manager at Labcon with a long history of working in the PCR industry, first as a chemist synthesizing oligonucleotide modifications, the critical reagent for PCR tests, and then as a dedicated technical support scientist. Utilizing his knowledge of dual-labeled probes, molecular beacons, scorpion probes, modified bases, fluorescent, and quenching modifications, he assisted Ph.D. level scientists on a daily basis in troubleshooting problems with qPCR and SNP assays. Over the years of working in this industry, Winsor has noticed common challenges that affect scientists working with this sensitive technology, and so SelectScience reached out to explore some of these shared hurdles in more detail.
What can go wrong and how can you tell?
During his time as a PCR tech support scientist, Winsor notes contamination was the number one cause of unexpected results. “Most of the time the contamination occurs at the site where the PCR is being performed. That’s why it’s important to ask yourself questions such as, is my work area decontaminated? Or, have I followed the proper decontamination protocol? When you have contamination, your test can be ruined by false positives or false negatives, sometimes even both, so it’s important to pay close attention to your controls.” Winsor goes on to detail that too great of an amplification or seeing amplification significantly earlier in your PCR runs could also be an indication of contamination.
Issues with experimental setup are another common hurdle for scientists working in the PCR lab, “if you don’t put in the right sequences, you will experience issues and won't be seeing amplification anywhere,” Winsor explains. “The way to get around that is to recheck your amplification. There's a lot of manufacturers that have really good software to check your primer and probes and whether you will be catching amplification based on your target sequence.”
There is also a chance that PCR probes and primers are received in too low of a concentration or too high of a concentration, and this can’t be detected without ultraviolet–visible spectroscopy unless you are using a dye-based product. Based on the molecular weight of your oligonucleotide, it should be possible to check whether you are working to the right specification. “Inconsistencies in concentration can happen at the PCR lab when someone has performed the wrong dilution, and it can happen at the manufacturing site because there was an error with pipetting or in the liquid handling software,” says Winsor.
What can I do if the issue is coming from the compounds supplied to me?
A pertinent source of irregular results in the PCR lab comes from inconsistencies in the compounds being used. Winsor explains that compound degradation due to shipping or ineffective packaging is a serious issue. “Compounds such as the probes used in the reaction are very sensitive. When these products are shipped over dry ice, without the right prevention measures in place, this can create an acidic environment, which can actually cause the fluorophore to fall off the oligo. Once this happens, there is nothing to tell you that amplification is occurring,” says Winsor.
5 key considerations for effective PCR
Decontaminate your working area
Ensure all your equipment is sterile
Double check all your concentrations
Verify the integrity of the compounds you are using
Check your DNA primers contain the right sequences
“The easiest way to tell if your compound is degraded is to look at your background fluorescence data. Most PCR machines and software will autocorrect a high background fluorescence data and effectively blank it out. If you are experiencing problems with high levels of background fluorescence, there is a chance that your oligo could have degraded, causing the quencher or fluorophore to have separated from the oligo, leaving the quencher fluorophore duo ineffective. Another way to check for oligo degradation is to run a HPLC analysis of your sample. If your fluorophore is no longer connected to your oligo chain, the retention time on your HPLC for fluorescence will be affected.”
Similarly, Winsor encourages scientists to use mass spectrometry to check that their sequences have been made in the correct order. “I have seen scientists struggling to achieve results only to realize that their DNA primer contained the wrong sequence. In this case, you can always ask the manufacturer to check their own quality control records.”
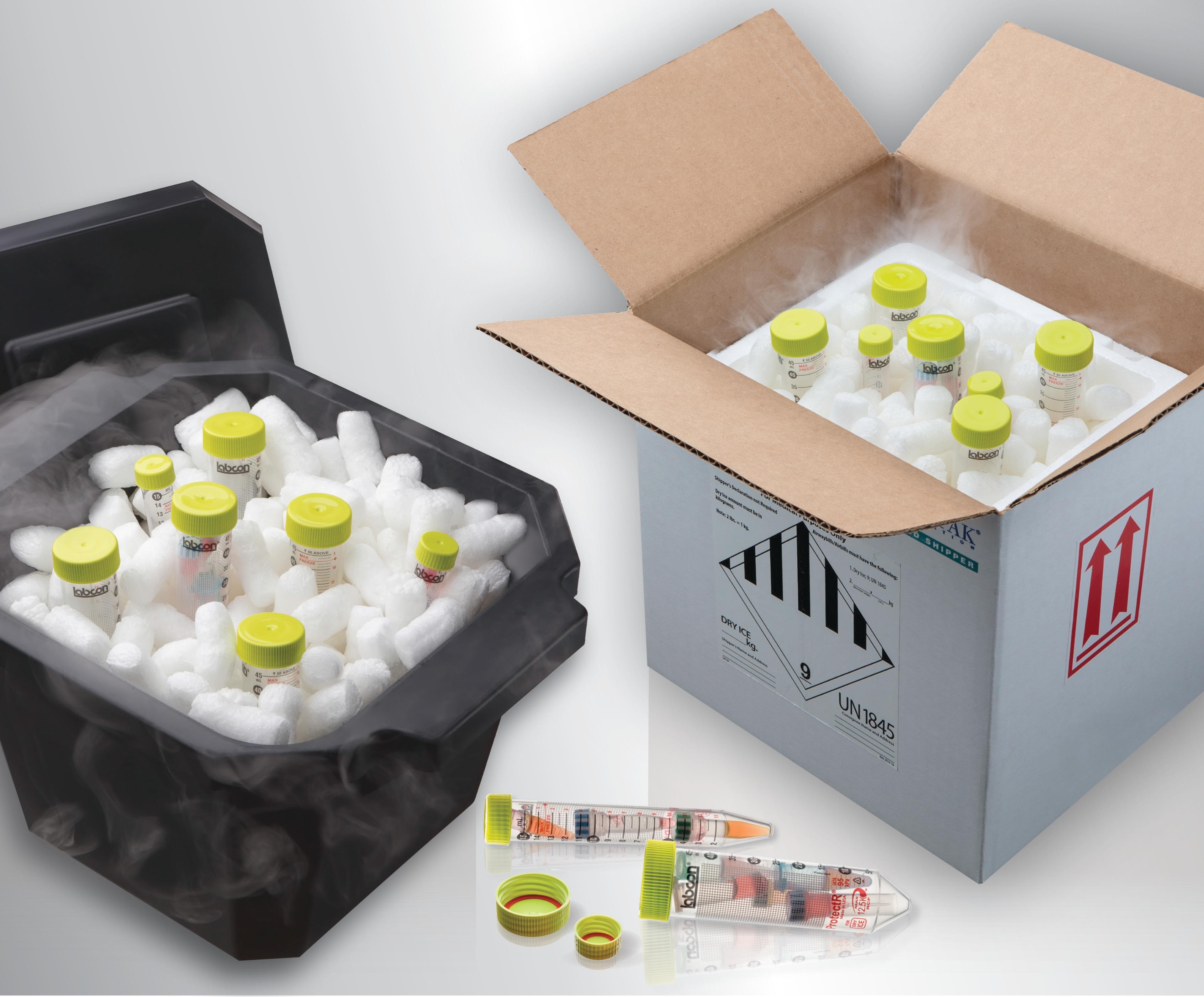
With the issue of acidification in mind, Winsor describes how Labcon developed its ProtectR™ Dry Ice Tubes, an IATA 95 kPa certified solution for manufacturers in this space offering full protection from pH damage caused by dry ice shipment. “The compounds used in PCR are a small but expensive material, so it is essential that every item they come into contact with is tested for impurities and have the appropriate certifications to go with them. All Labcon’s products undergo this level of testing to ensure our solutions are of the highest quality.”
Are there any questions I can ask my supplier in advance of placing an order to ensure quality?
To limit the likelihood of your reagents degrading, Winsor advises you should ask your supplier about their shipping and manufacturing processes, and what materials they're using for both. That includes anything that comes into contact with the reagents. “I would ask whether they are working in compliance with the industry recommended 10-6 sterility assurance level (SAL) when they're doing their liquid handling and their synthesis. When you're working with something like DNA, you want to maximize sterility at every turn,” he explains. “I would also ensure all materials and reagents are coming from an ISO 9001-certified facility. This means it has been verified by outside organizations and government entities with a view to safeguarding its customers.”