Impaired Protein Homeostasis in Neurodegenerative Diseases
Learn how selective antibodies help decipher an impaired autophagy pathway in spino-bulbar muscular atrophy
6 Mar 2018
Protein homeostasis, a state of balance in protein levels, is key to maintaining cellular health. In neurodegenerative diseases such as Alzheimer’s disease, as well as polyglutamine diseases such as spino-bulbar muscular atrophy, an impaired protein homeostasis triggers a physiological state where neurons are damaged. In this SelectScience® interview, Dr. Constanza Cortes, Assistant Professor in the Department of Neurology at Duke University School of Medicine, explains her research initiatives in studying proteostasis dysfunction and autophagy in neuromuscular and neurodegenerative diseases.
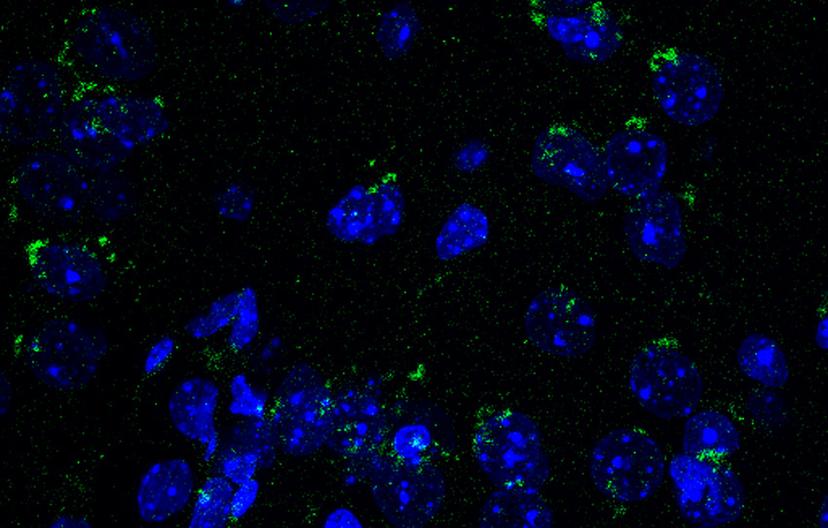
Mouse brain sections immunostained with BioLegend’s LC3 antibody. Image courtesy of the Cortes lab.

Dr. Constanza Cortes, Assistant Professor in the Department of Neurology at Duke University School of Medicine, studies impaired protein homeostasis in polyglutamine diseases such as spino-bulbar muscular atrophy.
SS: What are polyglutamine diseases?
CC: Polyglutamine diseases are a family of inherited neurodegenerative disorders, all caused by expansion of a triplet-nucleotide, CAG, in the coding region of the affected genes. The most notable is Huntington’s Disease, caused by a CAG expansion in the huntingtin gene. Another polyglutamine disease is spino-bulbar muscular atrophy (SBMA), caused by a CAG expansion in the androgen receptor (AR) gene. As the CAG codon encodes for the amino acid glutamine, the resulting proteins all carry extended polyglutamine tracts, giving the name to this family of diseases.
I study SBMA, a neuromuscular condition that targets the skeletal muscle and motor neurons in the lower spinal cord. These express high levels of the mutant AR, that result in protein inclusions, accumulating in the nuclei of affected tissues. This suggests that impaired protein quality control may form the basis of the neurodegenerative phenotypes observed in SBMA.
SS: What is the focus of your research in spino-bulbar muscular atrophy (SBMA)?
CC: I focus on protein quality control mechanisms and investigate the role they may play in the pathogenesis of SBMA. Protein homeostasis (also known as proteostasis), is fundamental for the survival of neurons, and proteostasis dysfunction is a feature of many neurodegenerative diseases, including Alzheimer’s and Parkinson’s disease. Understanding the mechanisms underlying proteostasis dysfunction and uncovering novel targets for rescuing these defects may yield important therapeutic targets for diseases associated with proteostasis failure.
SS: How does impaired protein homeostasis contribute to neurodegeneration in SBMA?
CC: I have uncovered a previously unreported periphery-to-CNS signaling network originating in skeletal muscle. Using a transgenic mouse model for a gene that’s a master regulator of cellular clearance and metabolism, conditionally expressed in the skeletal muscle, I have shown improved proteostasis in the CNS during normal aging. This suggests that maintaining skeletal muscle proteostasis during aging may yield important neuroprotective benefits in the aging brain.
In agreement with this, these muscle-specific proteostasis-activated mice perform significantly better in neurocognitive testing at 18 months of age compared to their age-matched control littermates. This suggests the existence of secreted signals originating in skeletal muscle and targeting the CNS, resulting in improved proteostasis control in the brain. My current work focuses on identifying those signals and testing their ability to rescue neurodegenerative ‘proteinopathies’, including Alzheimer’s disease.
SS: What are the methods you use to study autophagy in these mouse models?
CC: I am a cellular and molecular neuroscientist, and I use a variety of protocols to answer key questions regarding integrated physiology between the periphery and the CNS. Autophagy is a complex cellular recycling pathway with an orchestra-sized number of players. P62, one of the players in autophagy, is regarded as a classical autophagy cargo adaptor responsible for recognizing both protein aggregates and autophagosomal membranes. As tissues age, their autophagy activity declines, leading to the accumulation of protein aggregates which can be labeled using p62 antibodies. I use BioLegend’s p62 antibodies to determine the levels of protein aggregation in aging tissues as an indirect measure of autophagy function.
Similarly, LC3 is another classical autophagy marker that, upon cleavage, associates with early autophagosomal membranes and remains bound to autophagosomes until degradation in lysosomes. This makes it a powerful marker for autophagy activation and can be tracked both in western blotting (where the cleavage from immature LC3-I to mature LC3II isoforms is readily detectable by a change in migration) and in immunofluorescence of tissues and cells as distinct puncta that represent individual autophagosomes.
SS: Immunoblots and immunofluorescence form a significant part of your research. How do you choose your antibodies?
CC: Antibodies are essential tools for specific, targeted detection of specific proteins in tissues. Using BioLegend antibodies, I have been able to directly examine both the subcellular localization (via immunofluorescence) and abundance (via immunoblotting) of autophagy markers in a variety of tissues, including brain, skeletal muscle, liver and adipose tissue. Selectivity is one of the key factors I look for on any antibody products.
SS: You work closely with BioLegend on antibody development. Can you please share your experience?
CC: I am currently working closely with BioLegend to develop better, more specific antibodies that target autophagy and lysosomal markers. For example, there is currently no good antibody available that recognizes phosphorylated- transcription factor EB (TFEB), a fundamental tool to assess the activation status of this master regulator of autophagy. We are in talks to develop a phospho-specific antibody against Serine 142 of TFEB, a principal phosphorylation step in TFEB biology, and will be testing them both in immunoblotting and immunofluorescence assays.
Furthermore, my group is testing some newly developed pan-TFEB antibodies for immunofluorescence staining, using our novel conditional TFEB overexpressing mouse model. Recently, I have also become interested in neuroinflammation and its potential contribution to brain aging and neurodegenerative disease. I will be working with the immunology group at BioLegend to test some of their available reagents to study inflammation in the context of brain aging, and possibly develop novel assays to examine immune cells’ infiltration in the hippocampus and cortex during Alzheimer’s disease.
SS: What is the future of your research?
CC: The plasticity of the human brain, and its ability for self-repair and recovery has only recently been recognized. It is my goal to develop patient-relevant therapies for neurodegenerative disease, particularly for Alzheimer’s disease. Over 5 million Americans are currently living with Alzheimer’s disease, and although we have made important breakthroughs regarding our understanding of the basic biological processes leading to disease, there are no currently available therapies. My research aims to uncover novel therapeutic targets, uncovering new signaling pathways in the cutting-edge field of ‘integrated physiology’.
Do you use BioLegend antibodies? We'd love to hear from you! Please leave a review here.
