Collaboration brings clarity: Understanding SARS-CoV-2-host interactions
We speak with Dr. Ed Emmott, founder of The Emmott Lab & COVID-19 Mass Spec Coalition member, about his latest research into the disease
5 Oct 2020

As the race for an effective SARS-CoV-2 vaccine continues to attract attention both from the public and the pharmaceutical industry, it is important to recognize concurrent research efforts in further understanding viral-host interactions, as these could uncover new targets for therapeutics.
Dr. Ed Emmott is a researcher working at the University of Liverpool who aims to use the latest analytical technologies available to uncover how RNA viruses replicate and interact with the host cell. He set up ‘The Emmott Lab’ back in 2019 after pursuing the idea of combining his extensive experience in virology, stemming from a Ph.D. studying the coronaviruses, and his accumulated in-depth understanding of proteomics. The lab now forms an integral part of the university’s Centre for Proteome Research within the Department of Biochemistry and Systems Biology.
Since the creation of the lab, Emmott primarily worked on positive-sense RNA viruses, a grouping that shares features such as a genome which functions as an mRNA molecule that can be directly translated, and the deployment of protease enzymes during replication. More recently, Emmott has been interested not only in looking at the function of these proteases and their role in replication, but also in discerning whether they are “moonlighting in some other role,” such as shutting down the translation of host cell proteins or cleaving proteins involved in recognizing foreign viruses or non-self messages to inhibit the immune response. To support this work, the lab has also been developing single-cell proteomic methods and starting to apply this to virus research.
Collaboration during calamity
The COVID-19 pandemic has forced many in the scientific community to converge like never before, especially those who have a background working with the coronaviruses. Emmott is no exception to this, in fact he and his lab are helping to lead the charge as part of the COVID-19 Mass Spec Coalition (1). Emmott explains how the coalition was established as a “means to apply the expertise of the international mass spectrometry community to study the virus.” He tells SelectScience: “Our labs were empty. Our machines were lying unused. And the technology could be being put to use both in a translational sense to study patient responses, potentially learning new markers to stratify illness, but also in a basic biological sense to understand more about the virus.” Mass spectrometry has already played a crucial part in understanding SARS-CoV-2 infection, producing studies to elucidate the interactome or the phosphoproteome of affected cells.
Uncovering new targets means new therapeutic approaches
The Emmott Lab has recently published new research under preprint, titled ‘Characterisation of protease activity during SARS-CoV-2 infection identifies novel viral cleavage sites and cellular targets for drug repurposing’ (2). In this study, Emmott and his collaborators were able to expand on previous research identifying some of the proteases SARS-CoV-2 produces as targets for drug inhibition. As Emmott explains, here the focus of the study was to “try to understand what are the proteolytic cleavage events happening during infection and can we try to attribute these to viral or cellular proteases and work out which proteins are being cleaved.” Ultimately the aim is to clarify questions such as: “Which cellular proteins are being cleaved? Are these being cleaved in a way that benefits the virus? And is this something that we can make use of, potentially by drugging these proteins with pre-existing pharmaceuticals?” Emmott explains that the research was able to produce a shortlist of 14 protein substrates of the SARS-CoV-2 main and papain-like proteases and that, through a siRNA knockdown and inhibitor studies, they were able to produce a significant inhibitory effect on viral replication. The potential implications of these findings are not just apparent in understanding the mechanisms of proteolysis during viral infection, they could also provide new drug targets and ultimately new treatment strategies for the COVID-19 disease.
Miniaturizing high-containment experiments at lab capacity
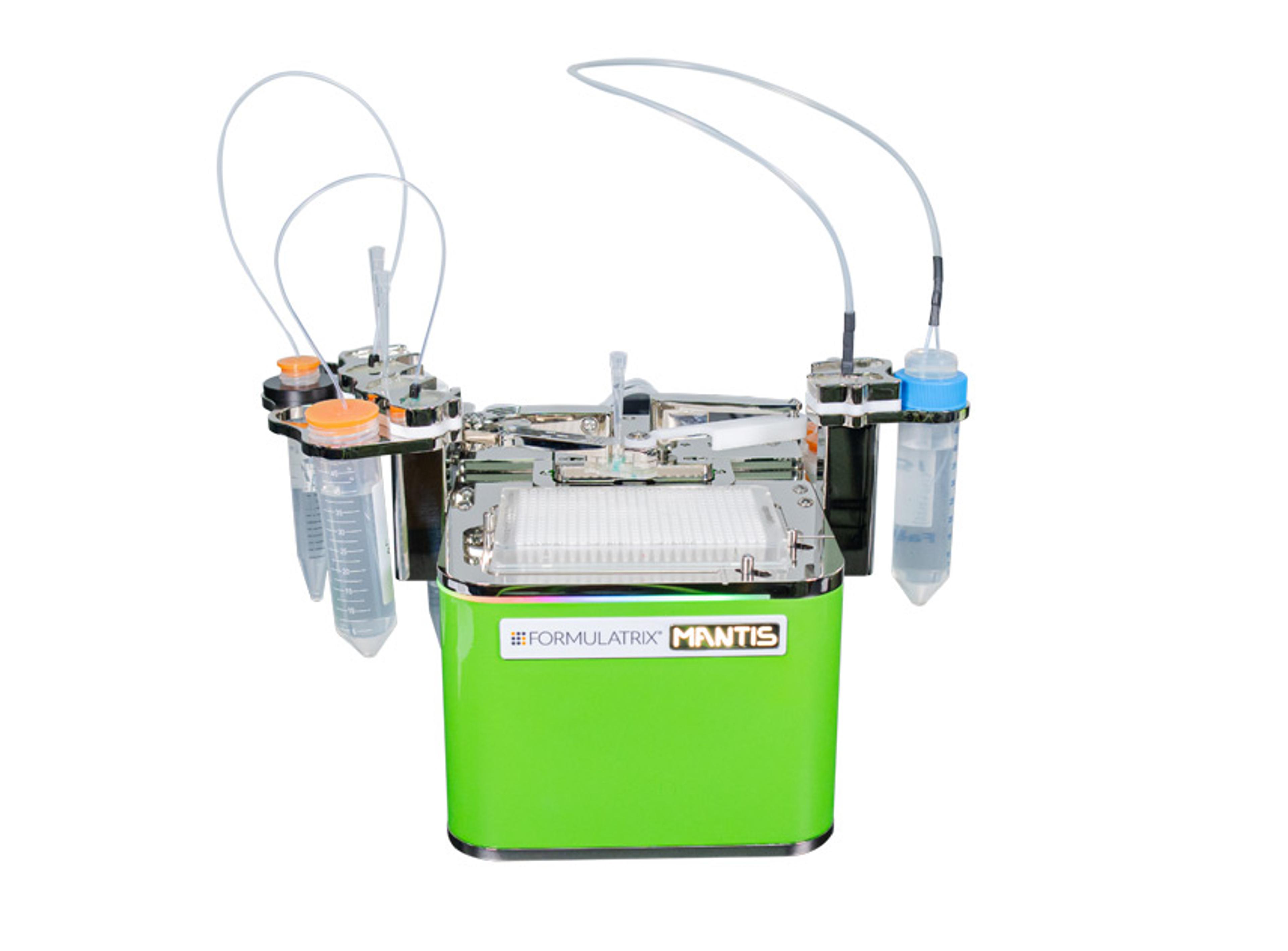
The MANTIS® Microfluidic Liquid Handler from Formulatrix
In order to perform its viral research, The Emmott lab heavily relies on automation and often high-throughput processing to meet the required number of experimental replicates. Their work is routinely done in plate-based formats using magnetic cleanup and multi-channel pipettors. However, for some projects, they do have to go to an even smaller scale. Emmott notes how the adoption of one particular system, the MANTIS® Microfluidic Liquid Handler from Formulatrix, has particularly benefited the lab’s work on SARS-CoV-2. He explains: “The MANTIS allows us to keep our sample volumes low, particularly when samples are very limited. This could be from working with previous patient samples or trying to minimize precious high-containment space and resources. Normally, you try to miniaturize sample processing to save reagent costs, and certainly that's nice, however in this case we actually get better data by miniaturizing as this lets us minimize loss of our limited material during handling. It's really a very good investment for our very low-input studies.”
Looking to the future
Emmott has been studying the coronaviruses throughout his career in viral research, but with the surge in investment, he has seen the basic science for SARS-CoV-2 progress rapidly. He believes high-throughput methods, such as mass spectrometry, can play a big role in this. However, with the host-pathogen interactions, Emmott admits, “there's still a lot to learn about the immune response to the virus. It's hugely important for discerning whether patients are responding well or not. Really, we need to understand that, especially in high-risk groups. It is going be particularly important for parts of the population who are less likely to respond well to a vaccine, for example, the elderly.”
Looking a little further ahead, Emmott sees cause for hope: “I'm optimistic that by the end of the year, we will have a vaccine licensed that will show some benefit against SARS-CoV-2, that people can start receiving early next year. I don't think it will be a perfect vaccine, it may only reduce very severe disease to a level where you feel ill for a few days. So, I think even with a vaccine that may be sufficient to allow life to return to a more normal setting, there will still be research ongoing to understand SARS-CoV-2 for years.”
Do you use Formulatrix products in your lab? Write a review today for your chance to win a $400 Amazon gift card>>
References
Struwe, W., Emmott, E., Bailey, M., et al. The COVID-19 MS Coalition—accelerating diagnostics, prognostics, and treatment. The Lancet; 395(10239):1761-1762 (2020).
Meyer, B., Chiaravalli, J., Brownridge, P., et al. Characterisation of protease activity during SARS-CoV-2 infection identifies novel viral cleavage sites and cellular targets for drug repurposing. bioRxiv. https://www.biorxiv.org/content/10.1101/2020.09.16.297945v1 (2020).

