Tracking COVID-19 ‘super spreading’ events with viral sequencing
Hear how one trailblazing lab at the Broad Institute of MIT and Harvard is using automated sample tracking to monitor pandemics such as Ebola Virus Disease and COVID-19
25 Aug 2020
Editorial article
The Sabeti Lab at the Broad Institute of MIT and Harvard is no stranger to handling difficult and dangerous viral outbreaks. During the Ebola Virus Disease (EVD) outbreak of 2013-2016, the lab was responsible for sequencing the Ebola virus (EBOV) genome, and their more recent work using single-cell gene expression analysis and transcriptomics of tissues in non-human primate models hopes to further aid the understanding of the pathology, progression, and evolution of EVD. However, with the emergence of the novel SARS-CoV-2 pathogen, the lab is now readjusting its focus and applying this existing expertise to address this new pandemic.
“The scale of this project was big, and it unfolded quickly. We had to rapidly build new systems to deal with this challenge,” Kat DeRuff, research associate in the Sabeti Lab, shares. “Luckily, we were able to adapt what we had set up for the Ebola project, and eLabJournal was absolutely crucial in this work.”
Genetic evidence for viral spread
Global genome tracing
The lab can also see how the spread in Boston relates to the rest of the world based on other research institutes that are uploading identified genomes. Through looking for links as to how these genomes connect, labs like the Sabeti lab can track how SARS-CoV-2 genomes are spreading and where they are spreading to.
The novel coronavirus is known to mutate approximately every 10 days, meaning that if a large group of people has the exact same virus, this must be spreading rapidly within that group without time to mutate. The main aim of the Sabeti lab’s project is to identify case introductions and dissect ‘super spreading’ events in the Greater Boston area. To achieve this, the lab is sequencing the SARS-CoV-2 genomes in patient samples and creating transmission trees based on who has what mutation.
How to align research and remote working
Three stages of sample processing
1. Biosamples received from collaborators with metadata in customized form
2. Biosamples labeled, scanned, and stored by the in-lab researchers
3. Metadata from collaborators matched up to the scanned biosamples by the end-user
Kat DeRuff was one of seven essential workers to continue this genome sequencing work in the lab once the pandemic hit. “If you look at the number of cases, it spiked really quickly,” DeRuff shares. “So, initially, we were getting hundreds of samples at a time and had few standardized protocols set up for processing and tracking these samples.”
However, once they had decided on how to standardize the labeling, storage, and tracking of samples, DeRuff notes that implementing this in eLabJournal from eLabNext was easy: “eLab made it so easy for us to organize and align our processes, even though we had different researchers, working at different times, and in different places.”

In an ideal research setting, one person would be responsible for the whole workflow. However, to keep in line with social distancing guidelines, researchers worked in two discreet teams in four-day rotations with one scientist working night shifts. With these new standards of working, the Sabeti lab split the work into two roles: in-lab user and end-user.
With the implementation of eLab, the integration of additional sample information could be done remotely by the end-user. Therefore, once the biosamples were received from external collaborators, in-lab researchers just had to make sure the samples were labeled correctly, scanned, and put into the appropriate freezer slot.
“At the beginning, this required a lot of back-and-forth, but once the original input was in place and we had our standardized system, the work could be done at any time by anyone who had access to the eLab software” DeRuff explained. “That was a really nice feature to have as it limited the amount of in-lab time that was spent on the computer and meant we could easily split up and coordinate those roles, especially when we weren't all in the same place.”
Streamlined, standardized, and automated protocols
The Sabeti lab received all its patient samples from collaborators at Mass General Hospital and the Massachusetts Department of Public Health. These collaborators would provide the biosamples along with corresponding de-identified data critical for interpreting the genetic sequence of the virus, such as sample collection date and volume of sample.
Working with collaborators meant that samples were received in many different formats, and in the beginning, the number of samples and the information provided was not standardized. DeRuff highlights that the real key to staying on top of this process was standardized labeling and sample metadata collection. “I think the first thing was making sure everything was labeled in exactly the same way,” DeRuff explains. “All you then needed to do is to scan this and store it in the correct freezer spot.”
DeRuff was also able to use eLab to create a specific metadata form for collaborators to help standardize the material they were receiving. Once this form was uploaded into eLab, the technology took care of the entire metadata tracking process. “We would take the original biosample, so the nasopharyngeal swab, and then extract the RNA to create ‘daughter’ samples for downstream analysis,” DeRuff explains. “eLab has a really great option where you only need to input the sample metadata for the original sample, which then passes automatically to the daughter samples when these are made.”

DeRuff credits eLabJournal as ‘critical’ to the streamlining and success of this work, highlighting the below benefits:
- “Label printing capabilities saved us a lot of time. We were handwriting these at first, but with eLab you can print customized labels for your entire set of samples. That was just a really, really great feature.”
- “We didn’t have consistent plate types and we were processing so many samples that we didn’t want to have to standardize that. So, one feature that worked really well was the ability to incorporate any barcode - it didn't have to be eLabNext-specific to be integrated into our system.”
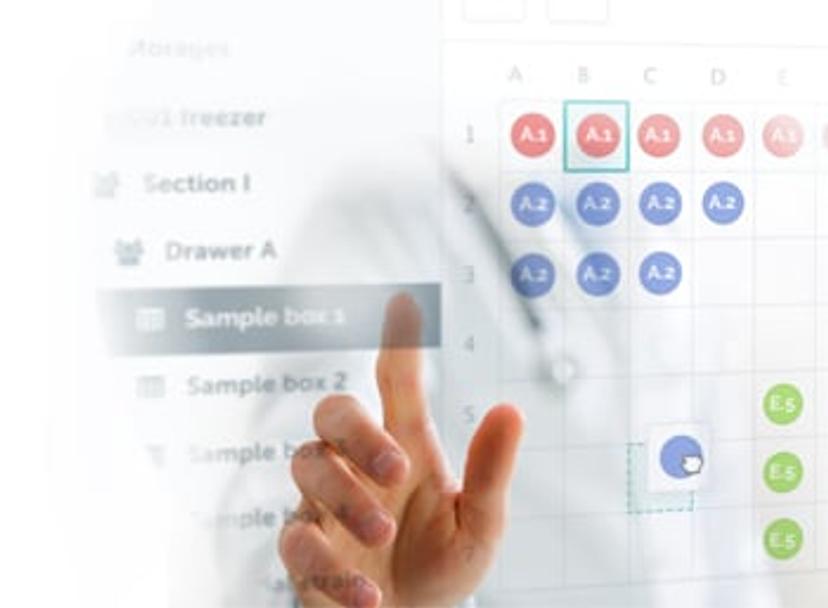
- "Another really helpful feature is the ability to electronically visualize your freezers. You can lay out a freezer in eLab to look exactly like your physical freezer, which made it really easy to see where something was digitally and easily find it in the lab.”
- "eLabNext were always so responsive in terms of helping us figure out how to create systems, print labels, and enable features to help us achieve what we needed. We wouldn't have been able to do this work without eLabNext’s technical support.”
Impacts of co-morbidities and public health decisions
As a continuation of its coronavirus research, the lab is using genomics to address gaps in research, including why some people continue to test positive for a long time, why others are not able to fight off infection, and how SARS-CoV-2 spreads beyond the lungs in severe cases. One of the main complications when reporting COVID-19 deaths is that there is no easy way to compare cases, due to the presence of co-morbidities. “I think what's making fighting this virus so challenging is that people who have co-morbidities are really vulnerable to it, but you don't actually know if patients are dying from the virus, dying from co-morbidities, or if it’s a combination, ” DeRuff explains.
The Sabeti Lab is contributing to the deployment of a contact tracing app that, when implemented with rapid diagnostics, hopes to make contact tracing easier.
Overall, DeRuff hopes that communicating these findings to those who dictate public health decisions will emphasize the need for robust early-detection systems and improvements to contact tracing protocols. “I'm really excited about how public health outcomes come from scientific research,” DeRuff explains, “The more information we have, the easier it is to elucidate the public health failures that happened, like the failure to both detect and control the circulating pathogen.”
Do you use eLabNext technology in your lab? Write a review today for your chance to win a $400 Amazon gift card >>


