Delve Deeper into Genetics with Revolutionary Dual CRISPR Technology
CRISPR Activation (CRISPRa) and CRISPR Interference (CRISPRi) offer precision, sensitivity and depth of understanding – find out more in this SelectScience expert interview
1 May 2019

Functional genomic screening links genetic information to phenotypic output, with the final goal of identifying and validating drug targets. Recent developments have seen two variants of the traditional CRISPR-Cas9 knockout system enable a deeper, more precise understanding of underlying genetic functions: CRISPR Activation (CRISPRa) and CRISPR Interference (CRISPRi). With CRISPRa as a gain-of-function and CRISPRi as a loss-of-function screening tool, researchers are able to design genomic screens to study both resistance and sensitivity to drugs.
In this article, we speak with Travis Hardcastle, associate product manager at Horizon Discovery, to get insights into the technology behind CRISPRa and CRISPRi, along with its applications, associated services and recent launch of the Dharmacon dCas9-VPR mRNA and synthetic SAM tracrRNA.
What are CRISPRa and CRISPRi and how do they modulate endogenous gene expression?
CRISPRa is a new method of artificial gene expression that utilizes a modified CRISPR gene editing platform to activate a gene of interest. This technique provides more specific expression than other techniques, such as using cDNA or ORFs, where there is often widespread overexpression that is not biologically relevant. CRISPRi offers a similar tool, but for gene knockdown.
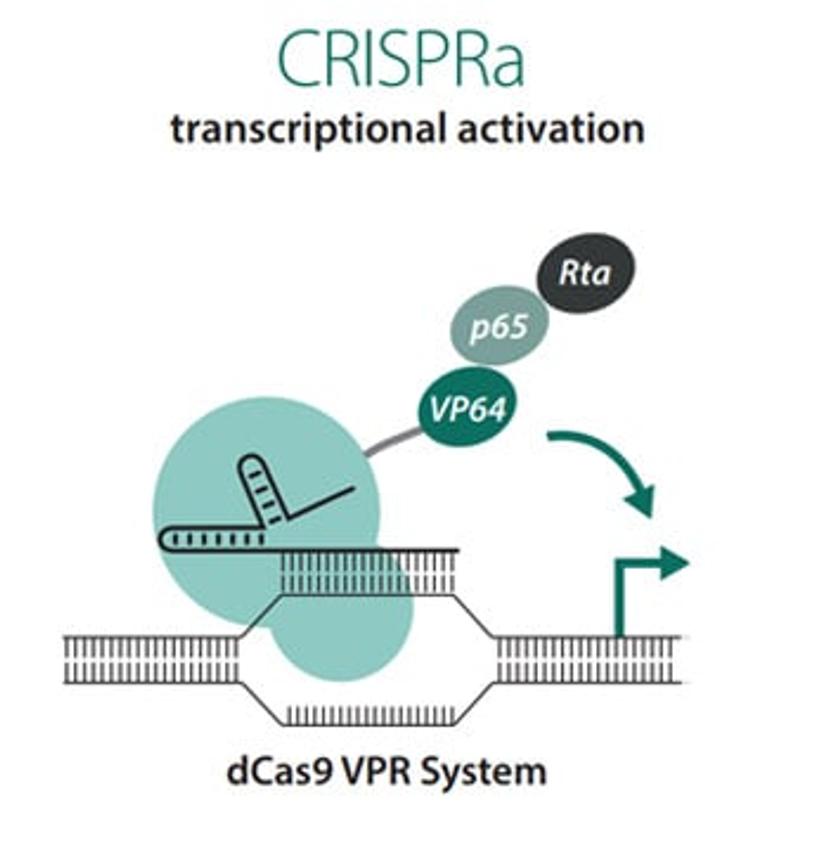
Our CRISPRa platform uses a deactivated Cas9 (dCas9), along with synthetic guide RNAs to target regions upstream of transcriptional start sites. The dCas9 is fused to three transcriptional activators, vp64, p65, and Rta, which comprise the VPR system, and when combined with gRNA leads to activation of the gene of interest.
We also recently launched our SAM tracrRNA for researchers using the Synergistic Activation Mediator (SAM) transcriptional activation system. This is the first commercially available synthetic tracrRNA option for SAM CRISPRa experiments. Both the VPR and SAM CRISPRa systems provide very robust endogenous gene activation.
The machinery is similar for CRISPRi, with the exception that dCas9 is fused to a KRAB domain that will inhibit transcription of the target gene.
What are the benefits of the CRISPRa approach?
CRISPRa offers a significant benefit over traditional overexpression methods, as it relies on the cell’s endogenous machinery to closely mimic native post-transcriptional processing. This allows researchers to express long or difficult-to-clone genes with CRISPRa, that cannot be expressed via traditional cDNA or ORF techniques. Also, CRISPRa allows the expression of non-protein coding genes, such as lncRNA. Finally, CRISPRa allows for the activation of multiple genes at the same time – which is another benefit you don’t traditionally see with cDNA or ORF technology.
I would encourage researchers to ask themselves, if you see a strong artificial activation, such as that seen with cDNA, how much biological relevance can you capture with that?
Travis Hardcastle Horizon Discovery
Horizon recently released a new dCas9-VPR mRNA option. How was that product developed and why?
Last year, we released our plasmid and lentiviral-based vectors for dCas9-VPR. During the development process, we learned that sometimes it's a challenge to develop a stable cell line. It takes a lot of time, several weeks, if not longer. Plus, some cell lines are just difficult to transfect.
So, our question was: can we develop a product that would offer the opportunity to perform quick one-off transcriptional activation experiments with something such as mRNA? If you want to investigate something in a one-off experiment, you can do this very quickly with dCas9-VPR mRNA. There is no need to develop a stable dCas9-VPR cell line. You simply transfect the cells, and 48 hours later the mRNA is gone. In a few days, you get a readout for what you were looking for – a phenotypic assay or RT-qPCR.
How is the CRISPRa/CRISPRi technology poised to revolutionize gene modulation screening?
One way I believe this technology is set to be revolutionary is through a dual screening approach, in which both CRISPRi and CRISPRa technologies are used in parallel. In combination with CRISPRi, a CRISPRa screen can reveal “switch-like” genes that display opposing effects when activated or inhibited in the presence of the drug of interest, bringing the prospect of unambiguous target discovery closer.
What are the benefits of dual CRISPR screening over the individual types of CRISPR screening?
CRISPR knockout screening is a great technique, but biology can be complicated and these knockout screens don’t always give us the whole picture. Dual screening can help reveal genes that behave differently when activated or inhibited in response to a drug of interest. This helps to identify new combinational therapy targets by identifying genes that promote either resistance or sensitivity. A CRISPRa and CRISPRi screen provides a vast amount of insight, enabling researchers to better understand interacting and opposing effects within the same signaling pathway.
Can you tell us about Horizon’s CRISPRa libraries?
Horizon’s libraries benefit from years of experience and our expertise. We have been doing this for a long time. In fact, Dharmacon released the first whole-genome siRNA reagent library for gene knockdown screening 15 years ago!
Horizon now also leverages that extensive expertise into our screening services, where we can help design the experiment, perform the assays and run the analysis to save the customer considerable time and frustration.
We focus on overarching themes of what plays an important role in biology and try to develop libraries that are associated with those pathways. Similar to our CRISPR knockout libraries, our CRISPR activation libraries are available for screening of gene families, druggable targets, and whole human genome. For example, we offer cell cycle regulation and epigenetic CRISPRa libraries, a cytokine receptor library with 110 gene targets, and a GPCR pathways library with 390 gene targets. We also have a fully customizable CRISPRa screening library option with our Cherry-pick Library Plater.
Our CRISPRa libraries are available in 96-well and 384-well plate format, targeting either human or mouse genomes. These plates are available as four individual crRNAs per gene or in pooled format, where all four crRNAs are pooled in the same well.
What’s the biggest story right now relevant to CRISPR screening?
I am probably a little biased on this, but for me it’s the fact that Horizon recently extended its CRISPR Screening Service to primary human T cells. This allows us to rapidly identify agents that modulate T cells, as single agents or in combination, at high throughput. In turn, we can provide analysis and interpretation of the effects of biologics or small molecules on multiple endpoints from multiple donors.
In your lifetime, what would you like to see science achieving?
I hope to see more progress in personalized medicine. If there is a single nucleotide polymorphism, can we correct it with gene editing? Is there a way in which we can harness personalized medicine to make it really personal for each individual? You know, that was one of the things that excited me the most about joining Horizon. The belief that we are providing cell-based solutions to help drive personalized medicine is at the core of all of our products and is really what Horizon aims to achieve.
Want to know more?
For more information on gene editing, screening and CRISPR technology:
- Watch the webinar: CRISPR screening in primary human T cells
- Read the article: CRISPR Screening for Genome-Wide Investigation of Gene Function
- Download the dual CRISPR screening poster
Does gene editing technology form a part of your work? We want to hear from you! Share your expertise.

