3D patient-derived cell models reflect tumor heterogeneity of clear cell renal cell carcinoma
Next-generation biobanking and cell culture technology for pathophysiologically relevant in vitro tumor models
1 Feb 2020

Unraveling the causes and consequences of tumor cell heterogeneity is a main research goal for Dr. Hella Bolck, senior research fellow in the Department of Pathology and Molecular Pathology at the University Hospital Zurich. Studying the remarkable molecular and phenotypic variability of tumor cells, including clinically important features such as their ability to seed metastases and to survive therapy, is a key focus. By considering the heterogeneity of cells within individual tumors, Bolck’s research aims to help develop better strategies to treat cancer.
Part of Bolck’s research requires the generation of patient-derived in vitro tumor models, which provide more pathophysiologically relevant cell culture tools for a variety of cancer biology projects. With a main focus on kidney cancers, a group of very heterogeneous tumors, the team at the department of pathology and molecular pathology want to understand whether patient-derived tumor models from clear cell renal cell carcinomas (ccRCCs) could reflect inter- and intra-patient heterogeneity in vitro. With the recent publication of promising results1, they now intend to dig even deeper in the longer run to provide pre-clinical cancer models from renal cancers and other tumor types that can mirror tumor heterogeneities to help understand their implications for cancer therapy. In this article, we talk to Dr. Bolck to find out more about 3D patient-derived cell models and their future potential.
What aspects of your work do you use 3D cell culture to focus on?
We are using 3D cell culture to generate pathophysiologically relevant cell models that can be used to study tumor biology and rationally design precision treatment strategies for patients. To do this, we have developed a next-generation comprehensive biobanking platform that includes the generation of patient-derived in vitro cell models alongside the diagnostic workflow at the department of pathology2. This is not only done for kidney cancers but also for a variety of other tumor types such a colorectal, pancreatic and lung cancer. As part of our biobanking strategy, we aim to generate 3D cell models that are representative of their respective cancers and can subsequently provide toolkits for many research projects addressing various questions in basic, translational and clinical research.

Why is 3D cell culture essential for your work?
Appropriate in vitro models are essential tools for cancer researchers. Immortalized cell lines that were used extensively in the past fail to reflect several important features of human tumors, for example, their subclonal architecture and phenotypic complexity. To achieve a more adequate representation of cancer in vitro, we are culturing patient-derived tumor cells. The idea is that such models are directly from the patient and thus very closely resemble the human tumor. 3D cellular model systems are a particularly attractive approach for this because they promise to mirror the in vivo tissue environment better than monolayers. There have been many prominent studies highlighting the potentials of 3D cell culture and so we use this methodology to develop more relevant cancer models.
How can 3D cell culture complement traditional cell cultivation in two dimensions?
3D cell cultures offer the possibility to model certain aspects of tissue architecture and reflect cell-cell and cell-environment interactions better than 2D cultures. These features are particularly important to study processes in which tissue organization or functional cooperation between cells affects pathophysiologically relevant processes, such as cell differentiation, proliferation or dissemination.
Monolayer cell cultures lack this spatial complexity but have offered important insights into tumor biology and are certainly still an important model system for cancer research. To derive kidney cancer models, we continue to rely on 2D models because they are robust and amenable to many downstream techniques. However, the additional layer of complexity and unique properties of 3D organoid cultures open up new possibilities for translational observations. We routinely use Corning® Matrigel® Growth Factor Reduced (GFR) Basement Membrane Matrix, Phenol Red-free, to generate our 3D tumor organoids. For us, the most appropriate strategy is to generate 3D tumor organoids and monolayer cultures in parallel because after all, the question, “which model system is most suitable?”, depends on the research goals in a particular project.
Which key results has your laboratory found by using this technology?
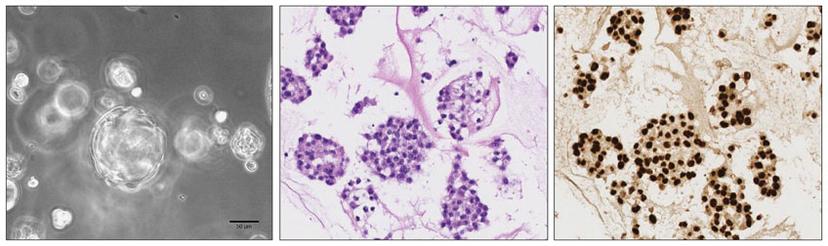
We have successfully generated 2D and 3D patient-derived cell models from clear cell renal cell carcinomas (ccRCCs) and confirmed their concordance with the original tumor tissue. We then addressed the ability of modeling ccRCC subclonal architecture in vitro and observed heterogeneous cell populations in both 2D monolayer and 3D organoid cell cultures. In fact, both cell culture models showed a high degree of similarity with the subclonal composition of the corresponding parental ccRCCs, which indicated that patient-derived kidney cancer models are able to represent tumor heterogeneity of ccRCCs in vitro. Moreover, we could detect clonal dynamics during serial passaging of cells, suggesting that patient-derived cell cultures may offer insights into evolutionary potential and treatment susceptibility of ccRCC subclones in vivo.
In our proof-of-principle drug response profiling, we found patient-specific sensitivities to clinically approved ccRCC therapeutics that could not be anticipated by interrogating commercially available cell lines. This has important implications for advancing precision cancer treatments and underscores the importance of developing relevant patient-specific cancer models such as tumor organoids.
What do you see as the future for 3D cell culture and model systems?
There have been many interesting developments in the field, for example, different 3D co-culture, microfluidics or organs-on-chips systems and it is great to see that the community is striving to make use of these tools to answer biological questions. I am, for example, very excited to read the first studies reporting clinical relevance of patient-derived 3D cell model systems for predicting drug responses and I hope there will be more like this to come. Besides these success stories, I think it is important to keep in mind that the effective analysis of complex cell systems such as 3D models can be challenging, and for many newly developed systems and associated technologies appropriate guidelines and best practices are not yet determined. It will be important to develop an awareness of the strengths and limitations of emerging systems and I hope that we will also see further studies determining the relevance and value of different cell culture tools in order to use them to their full potential.
References
Bolck HA, Corrò C, Kahraman A, von Teichman A, Toussaint NC, Kuipers J, Chiovaro F, Koelzer VH, Pauli C, Moritz W, Bode PK, Rechsteiner M, Beerenwinkel N, Schraml P, Moch H. Tracing Clonal Dynamics Reveals that Two- and Three-dimensional Patient-derived Cell Models Capture Tumor Heterogeneity of Clear Cell Renal Cell Carcinoma. Eur Urol Focus. 2019 Jun 29.
Bolck Hella A., Pauli Chantal, Göbel Elisabeth, Mühlbauer Katharina, Dettwiler Susanne, Moch Holger, Schraml Peter. Cancer Sample Biobanking at the Next Level: Combining Tissue With Living Cell Repositories to Promote Precision Medicine. Frontiers in Cell and Developmental Biology. 2019 Oct 22.
