Nucleic Acid Metrology: The Value of Inter-laboratory Reproducibility in Digital PCR
Dr Jim Huggett, LGC & University of Surrey, discusses the results of a study into international inter-laboratory digital PCR
14 Mar 2017

Dr Jim Huggett, Principal Scientist at LGC, UK LGC LGC is a global leader in the laboratory services, measurement standards, reference materials, genomics and proficiency testing market places.
A recently published inter-laboratory digital PCR study, demonstrated high reproducibility for the measurement of rare sequence variants, using digital PCR. SelectScience spoke to the paper’s senior author, Dr Jim Huggett, to learn more.
SN: Could you confirm your name, full job title and place of work for us?
JH: My name is Dr Jim Huggett and I am a Principal Scientist at LGC. I am also a Senior Lecturer at the University of Surrey, School of Biosciences and Managing Editor of the Journal, Biomolecular Detection and Quantification.
SN: What are your main job responsibilities?
JH: I coordinate a team of scientists in a research field applying metrology to molecular biology. Metrology is the science of measurements, which is well established in fields such as physics and chemistry, but in biology it is a relatively new concept. Metrology enables us to investigate the accuracy of a method and how to improve this, as well as to develop mechanisms for ensuring different laboratories are able to make comparable measurements. We study a wide range of biological measurements applied to a variety of scenarios including RNA biomarkers, cancer genotyping and microbial analysis.
Nucleic acid metrology in the clinical and preclinical arena is challenging for many reasons, including the fact that a biological system, by its very nature is variable. In molecular biology, it is extremely important that methods are robust, however the technical error of the methods is usually not considered.
The value of digital PCR
SN: Could you explain Droplet Digital PCR technology to us, how does it differ from traditional PCR techniques?
JH: PCR can be split into traditional, real time and digital. Traditional PCR allows us to detect a DNA sequence by amplifying it and assessing it at the end of the reaction, using another method like agarose gel electrophoresis. Real-time quantitative PCR (qPCR) allows us to measure the accumulated amplified product, during the reaction, in ‘real-time’. This is able to provide a precise estimation of the amount of DNA so it can be used for quantification, such as when measuring the amount of virus in a hepatitis B patient.
While the method is precise, making reproducible measurements between laboratories can be very challenging, because the method is a relative, analogue approach; the output quantification cycle (Cq, also termed cycle threshold or Ct) can vary when measuring the same sample for a variety of reasons. Standardization approaches aligning to endogenous controls or calibration curves are used, but employing them is very challenging. Consequently qPCR remains a precise but frequently biased method, leading to poor reproducibility when used on biological samples. This may explain why qPCR is used extensively in biomarker research, but few of these biomarkers are translated to aid patient care, and its use in routine diagnosis is comparatively rare.
Digital PCR (dPCR) is a binary PCR technology achieved by conducting a limiting dilution of the DNA molecules across hundreds, thousands, or even millions of tiny reactions called partitions. The term droplet in this context refers to a method for partitioning the reaction using water-oil droplets. Following PCR amplification, the positive and negative partitions are counted as a measure of those that contain DNA or not, respectively. This provides an absolute quantitative value so that dPCR results are less ambiguous, do not rely on a calibration curve, and can provide better precision, sensitivity and specificity than qPCR.
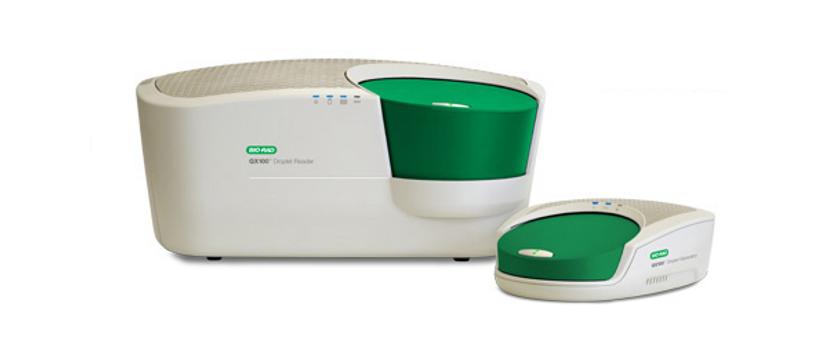
QX100™ Droplet Digital™ PCR System
Overcoming the obstacles of qPCR
SN: You recently published a paper on inter-laboratory dPCR reproducibility, can you tell us why there is a need for high reproducibility PCR?
JH: One of the biggest obstacles to the adoption of quantitative molecular methods in the clinical arena has been the inability of the methods to perform with high reproducibility; in other words it is not that easy for you and I to get comparable results. Reproducibility is extremely important for clinical and preclinical research, and lack of reproducibility is a big disadvantage of qPCR in the laboratory.
The binary and absolute quantification of nucleic acid means that dPCR should provide higher inter-laboratory reproducibility than qPCR. This is what we set out to test in our recent study.
We asked 21 participating laboratories to test for a KRAS point mutation, one of the most challenging types of cancer biomarkers to identify. We prepared all of the test materials at LGC (UK) and the samples were analyzed at the participating laboratories, using the BioRad QX200TM or the BioRad QX100TM Droplet Digital PCR system. Three levels of mutation concentration were analyzed in a background of wild type sequence, from ~0.2% to 12% fractional abundances. The participants in the study had various levels of experience with dPCR, ranging from being highly experienced, to being relatively new to the technique.
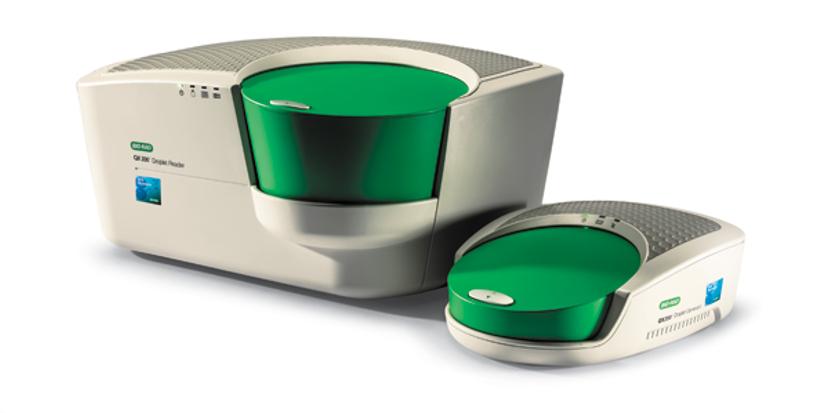

QX200™ Droplet Digital™ PCR System
SN: What were the findings of the study?
JH: Astonishingly, all of the participating laboratories achieved the same results, and we found no statistically relevant differences in the results. Initially, three of the laboratories were out on the mutant sequence, but these results were corrected following the application of follow-up guidance.
Our results demonstrated that fractional abundancies and absolute quantities of mutant sequence, when present as low as 0.2%, can be reproducibly quantified using dPCR, without the need for calibration. This is simply not possible using qPCR, or for that matter, with next generation sequencing (NGS).
SN: What is the significance of these results?
JH: Discussions around dPCR have often focused upon the methods ability to measure rare mutant targets, quantify with high precision or measure genetic relationships where the technique has an advantage over other methods. Such application offers potentially new biomarkers to be measured. What we show here is unprecedented reproducibility, the labs were able to get comparable results with little effort; they simply counted the DNA molecules. This characteristic, while perhaps less exciting than accessing a new biomarker, is potentially dPCRs most powerful accolade. All of a sudden it is much easier for you and I to get the same result; this could have huge potential in clinical research and diagnosis.
In the short term, I see dPCR being used as a reference method, to improve harmonization between laboratories using other method like qPCR, for example when testing viral loads. dPCR could be used to reduce variability between testing laboratories, and in clinical research, scientists may find that they are able to get more comparable results. I’m sometimes asked if dPCR will replace qPCR, and the answer to that is that while this is possible, it is unlikely to happen anytime soon. While dPCR has obvious reproducibility advantages, qPCR is well established, high-throughput and is valuable to clinical manufacturers. While dPCR is very powerful and based on the same underlying chemistries as qPCR, the technique still requires a lot of development.
But dPCR has many possible future applications in areas such as in supporting cancer diagnosis and treatment, but it also has potential in the point-of-care market. In near patient testing, the automated quantification of nucleic acids for rapid diagnostics has much potential, and the simplified binary readout of dPCR lends itself to automation far more than the qPCR amplification curve. It is a very exciting time to be working in the molecular diagnostics field and I suspect dPCR will have an important role to play in advancing human health.