Application Note
Resources
25
Selected Filters:
White Papers
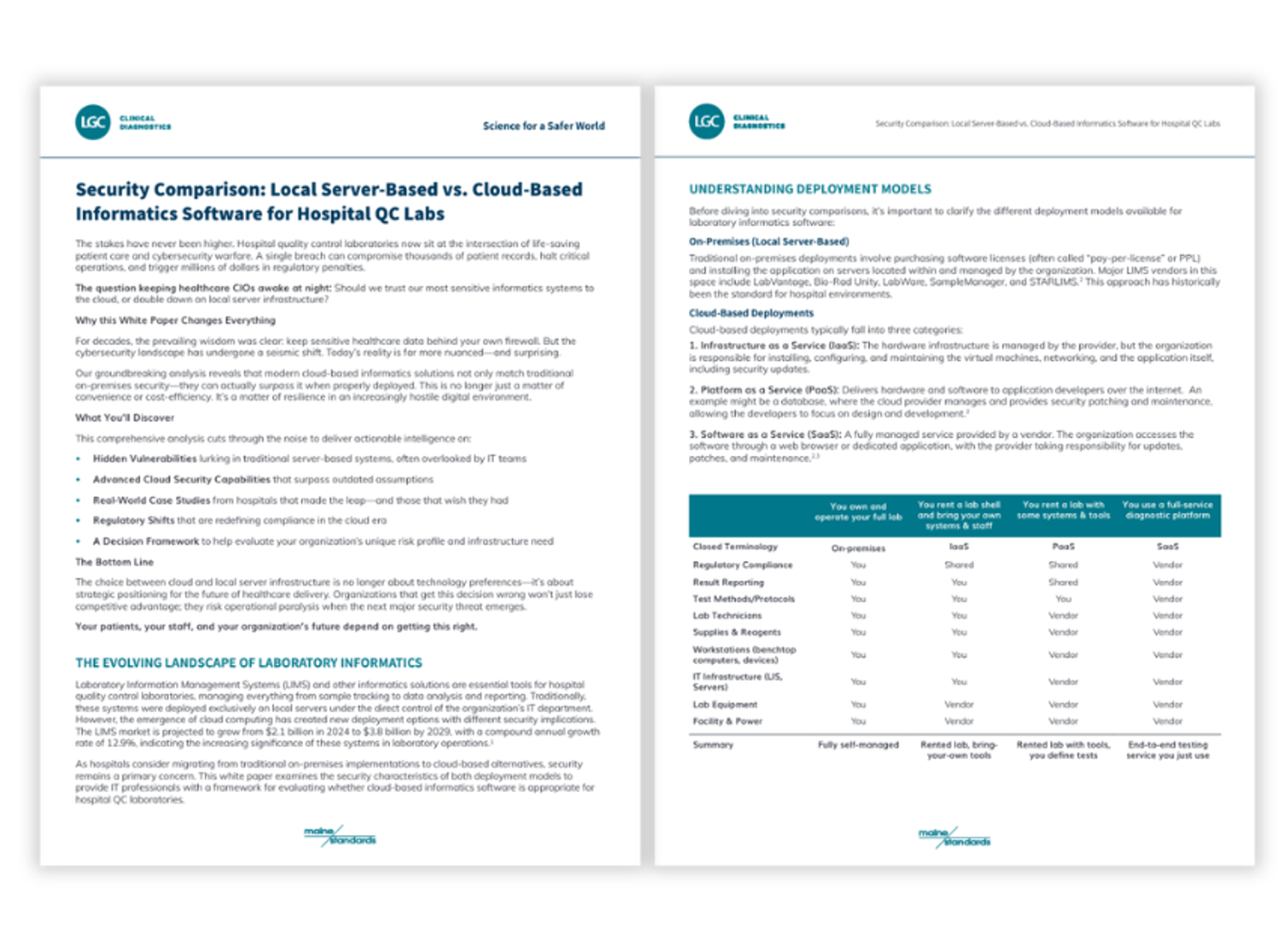
Turning atoms and lasers into a quantum computer
Application Note
DRAGEN secondary analysis
Product Brochures
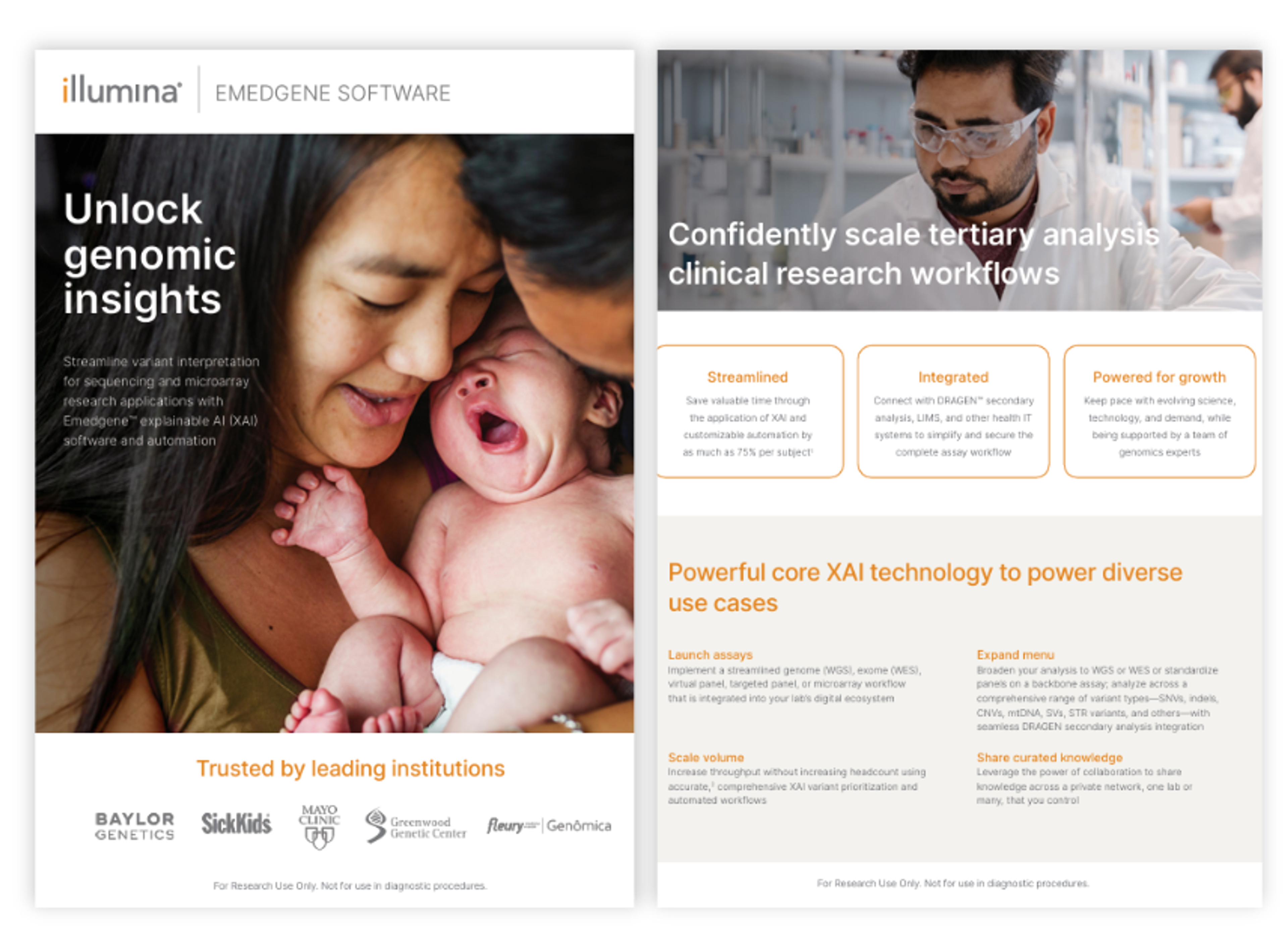
Unlock genomic insights with Emedgene software
Application Note
Emedgene software
White Papers
Accelerating science and driving productivity
White Papers
The concept of 21 CFR Part 11 without a PC
Product Brochures
Accelerate your lab performance with CrossLab Connect
Product Brochures
Your all-in-one electronic lab notebook, LIMS & informatics solution
Product Brochures
Manage and share analytical data with context
Product Brochures
Chemical structure drawing software
Product Brochures
Method Selection Suite: Robust methods with fewer injections
White Papers
Why NGS ELN deployments fail
Product Brochures
Industry standard property calculators
Product Brochures
Deformulate and identify MS components
Product Brochures
Percepta flyer
Product Brochures